Introduction
Dongtingking crucian carp, Carassius auratus indigentiaus is a new subspecies of crucian discovered by our research team in Beimin Lake, Li County, Hunan Province. It is one of the unique species remaining in the marginal lake water area during the historical change process of Dongting Lake.1 Dongtingking crucian carp exhibits characteristics such as rapid growth (over 10% faster than the widely cultivated Pengze crucian carp nationwide), high nutritional value (protein content 18.63%, amino acids 866.12 mg/g dry weight), delicious flavor (umami amino acids accounting for up to 50%), stable genetic traits, pure germplasm, and a high recapture rate. Farmed in the Yangtze River and Pearl River basins, it is highly favored by consumers. It has already generated economic benefits exceeding hundreds of millions of Chinese Yuan (CNY), indicating very broad prospects for further development of its aquaculture.2
However, under high-density farming conditions, Dongtingking crucian carp are susceptible to various diseases, and during fry hatching, fish eggs are prone to fungal infections, leading to saprolegniasis. Additionally, they may suffer from gill rot disease caused by bacteria or viruses.3 These disease threats pose a major constraint to the sustainable expansion of its aquaculture, necessitating a deeper understanding of its innate defense mechanisms to develop effective prevention strategies.
The skin of fish is directly contacted with water environment which is covered with a layer of mucus secreted by epithelial cells and epidermal goblet cells. As the first immune defense line of fish, mucus fulfills essential physiological functions including osmoregulation, lubrication, facilitation of intraspecific chemical communication, and immune activity.4 The mucus is rich in diverse immune-related proteins (e.g., immunoglobulins, lectins, lysozymes, complement, antiviral protein, antimicrobial peptide, SOD enzyme) that directly combat pathogens, prevent colonization, and modulate inflammatory responses.5 There are sporadic reports on genetics, development, and immunity of Dongtingking crucian carp,6–8 Proteomics has been involved in aquaculture in recent years, but the specific composition and immune-related protein profile of its skin mucus proteome remain entirely unexplored. This gap hinders a molecular-level understanding of its disease resistance mechanisms.
To gain an in-depth understanding of the molecular composition and immune-related proteins of skin mucus in Dongtingking crucian carp, we employed a proteomic approach for the first time to analyze the mucus proteome. Mucus proteins were extracted, reduced alkylated and enzymatically hydrolyzed, and 1903 kinds of proteins were identified by proteomics strategy. GO analysis and KEGG related function enrichment analysis were carried out for the above proteins. This study laid a foundation for understanding the protein molecular diversity and immune mechanism of Dongtingking crucian carp mucus.
Materials and Methods
Mucus collection
Dongtingking crucian carp were collected from South Aquatic Technology Innovation and Training Base in Hunan University of Arts and Sciences. Five healthy Dongtingking crucian carp with similar body weights (approximately 250 g each) were selected. The Dongtingking crucian carp were buried in ice for 5 minutes to induce unconsciousness for subsequent mucus collection. A single collection bag, pre-filled with 1 mL of physiological saline, was used for all fish sequentially. Each fish was individually placed into the same bag in a dorsal (back-down) position to minimize contamination from excretions. Care was taken to ensure that only the dorsal skin contacted the saline, thereby avoiding contamination from gill mucus and waste. The dorsal skin was then gently kneaded and stimulated to induce mucus secretion. To ensure sufficient yield, residual mucus was further scraped from the dorsal skin using a plastic sheet. After processing each fish, the accumulated mucus in the saline was temporarily held in the bag. After all fish were processed, the total pooled mucus from the bag was aspirated with a pipette, transferred into 2 mL centrifuge tubes, and stored at -80 ℃.
Sample preparation
Mucus sample stored at -80 ℃ was added with lysis solution [containing 1.5% SDS and 100 mM Tris-HCl (pH 8.0)] at a 1:10 mass/volume ratio. The sample was homogenized at high speed twice for 1 minute each time, and then sonicated by using ultrasonic cell disintegrator (Ningbo Xinzhi JY92-ⅡN) for 15 minutes (2#rod, power 20%). Subsequently, sample was centrifuged at 12,000 g for 15 minutes at room temperature, and supernatant was collected; Next, four times the volume of precooled acetone at -20 ℃ was added, mixed thoroughly, and then the supernatant was allowed to stand overnight at 4 ℃. The mucus sample was centrifuged at 12,000 g for 15 minutes at 4 ℃, and then supernatant was discarded. Pellet was washed three times using an appropriate volume of pre-chilled acetone by centrifugation (12,000 g, 15 minutes, 4 ℃) followed by air-dried naturally. Protein samples were reconstituted in lysis solution (8 M urea prepared in 100 mM Tris-HCl buffer, pH 8.0), with the addition of dithiothreitol (DTT) to a final concentration of 10 mM. The mixture was then incubated at 37 ℃ for 1 hour to facilitate the reduction reaction and open disulfide bond. Subsequently, iodoacetamide (IAA) was added to a final concentration of 40 mM for the alkylation reaction, which was performed at room temperature in darkness. Protein concentration was measured using the Bradford method with a kit from Sangon Biotech (Shanghai). The reduced and alkylated sample, was then mixed with a 100 mM Tris-HCl solution (pH 8.0), and the concentration of urea was reduced to below than 2 M. Trypsin was added at an enzyme-to-protein mass ratio of 1:50, and the enzymatic digestion was carried out by overnight incubation at 37 ℃ with agitation. The following day, trifluoroacetic acid (TFA) was introduced to halt the enzymatic digestion, and the solution’s pH value was adjusted to approximately 6. The supernatant was centrifuged at 12,000 g for 15 minutes, and the resulting supernatant was collected for desalting using a Stage Tip. The peptide segment solution after salt removal was dried using a centrifugal concentrator and stored frozen at -20 ℃ for future use.
MS and protein identification
Mass spectrometric analysis was conducted on a Q Exactive HF-X hybrid quadrupole-Orbitrap mass spectrometer (Thermo Scientific). Peptide mixtures were automatically loaded onto C18 trap column via an autosampler, followed by eluted onto an analytical column (75 μm × 250 mm, 2 μm particle size, 100 Å pore size, Acclaim PepMap C18 column, Thermo) for separation. Two mobile phases (mobile phase A: 0.1% formic acid; mobile phase B: 90% ACN, 0.1% formic acid) were used to establish a 120-minute analytical gradient with the following profile: 0 min at 5% B, 80 min from 5% to 23% B, 20 min from 23% to 52% B, 5 min from 52% to 80% B, 80% B for 5 min, 0.1 min from 80% to 5% B, and 5% B for 9.9 min, at a constant flow rate of 300 nL/min. In the MS DDA mode analysis, each scan cycle comprised a one full MS scan (R=60 K, max IT=50 ms, scan range 300–1800 m/z) succeeded by 20 MS/MS scans (R=15 K, max IT=110 ms). The collision energy for HCD was set at 28, and the quadrupole isolation window was defined as 1.2 Da. Dynamic exclusion for repeated ion acquisition was set at 30 seconds. After mass spectrometry detection was completed, the obtained data required database search to reveal the final protein identification results. In this project, the database Carassius auratus.fasta was used for the search. The raw mass spectrometry data, (stored in raw format), contained complete scan information. The acquired raw file was then directly imported into the software named Proteome Discoverer 2.3 for database search, as well as for peptide spectrum matching and protein quantification. The analysis parameters for Proteome Discoverer 2.3 are provided in Table 1. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository9,10 with the dataset identifier PXD071967.
Results
Protein Identification Results
Proteins from the mucus of Dongtingking crucian carp were successfully identified via mass spectrometric database searching. The data were organized using Microsoft Excel, resulting in the identification of 1,903 proteins (Table S1). Information on selected top-ranking proteins is presented in Table 2.
Overall Analysis of Mucus Proteins in Dongtingking Crucian Carp
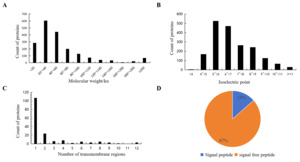
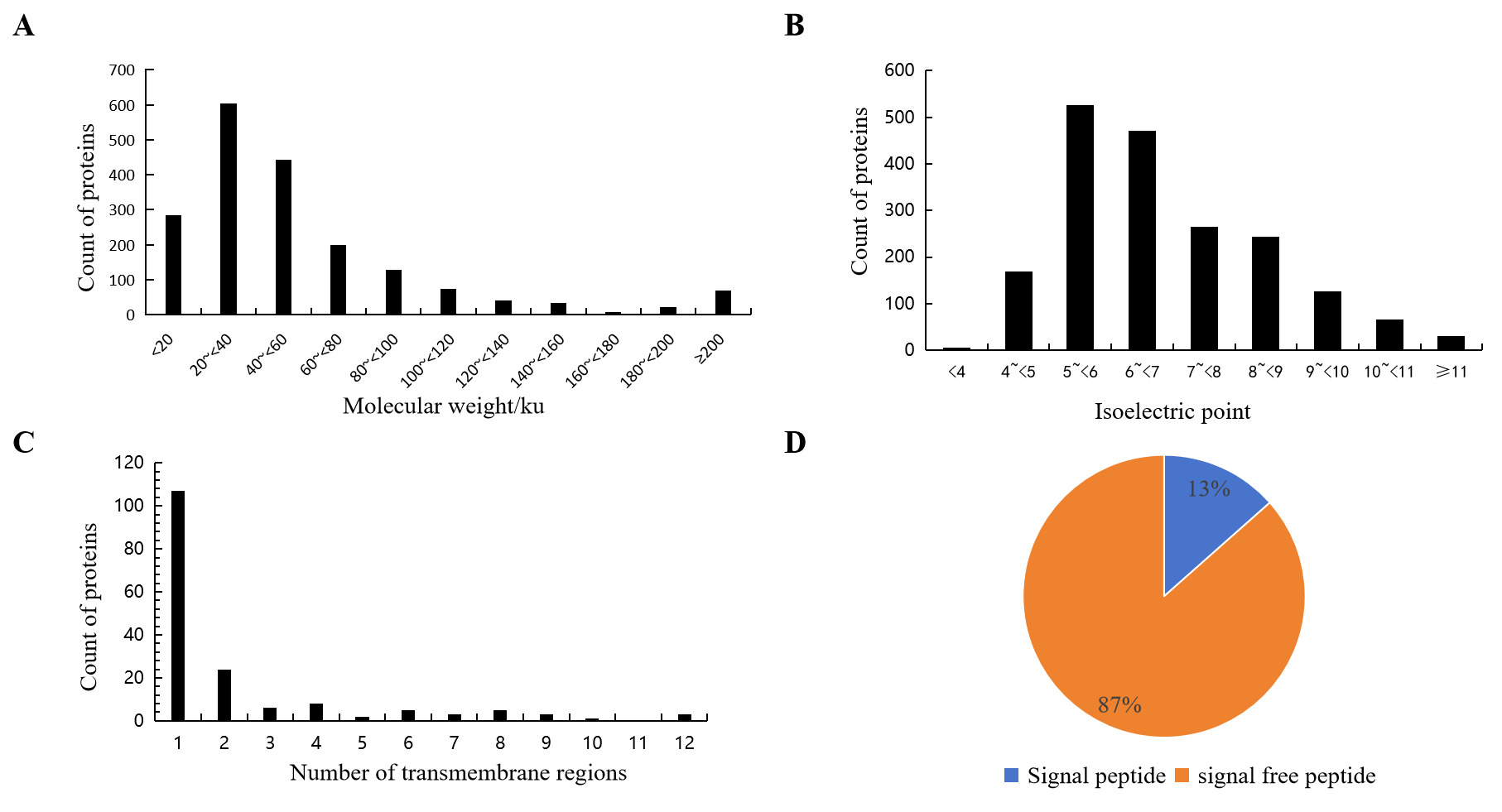
The molecular weight of mucus proteins in Dongtingking crucian carp ranged from 4.9 to 1004.5 kDa. Among them, 1,656 proteins (87.0%) exhibited molecular masses between 4.9 and 100 kDa (Fig 1 A). The isoelectric points (pI) of these proteins spanned from 3.4 to 12.2, with 998 proteins (52.4%) showing pI values clustered between 5 and 7 (Fig 1 B). Prediction using TMHMM-2.0 indicated that 167 proteins possessed at least one transmembrane domain (Fig 1 C). SignalP 5.0 prediction revealed that 256 proteins (13%) contained signal peptide sequences (Fig 1 D) including vitellogenin(A0A6P6J2V1), macroglobulin(A0A6P6QXH9), calreticulin(A0A6P6J056), cadherin(A0A6P6PUW9), calnexin(A0A6P6QQ28), cathepsin(A0A6P6R2K6), collagen(A0A6P6JJT0), complement(A0A6P6K562), lectin(A0A6P6MTU9), mucin(A0A6P6MDT6), among others (Table S1), suggesting that these proteins are secretory proteins.
GO Enrichment Analysis
GO enrichment analysis was conducted for the successfully identified proteins from skin mucus. Regarding molecular function, proteins with binding functions were the most abundant, comprising 1,435 entities, followed by those with catalytic activity, numbering 717. In terms of cellular component, intracellular anatomical structure constituted the largest group of proteins, followed by cytoplasm and organelle. For biological process, the proteins were predominantly enriched in the following processes: metabolic processes, cellular component organization or biogenesis, macromolecule localization, cellular localization, and response to stress, etc. (Figure 2).
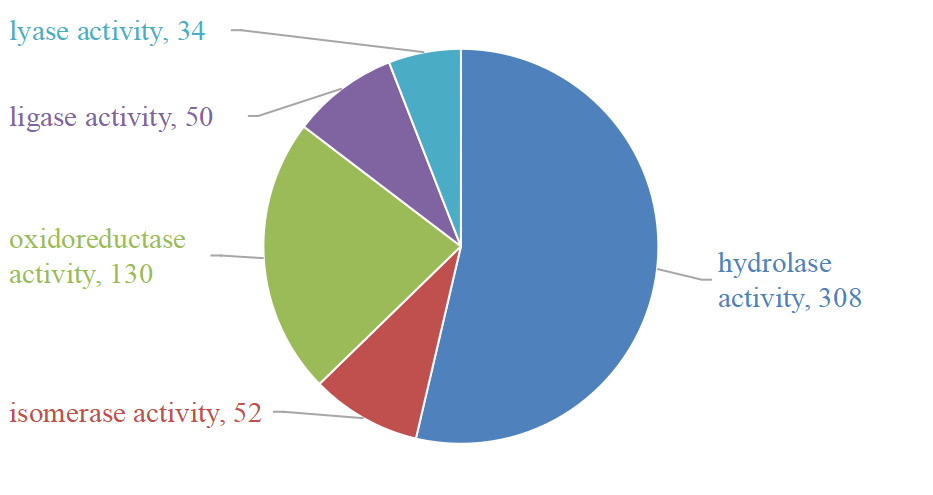
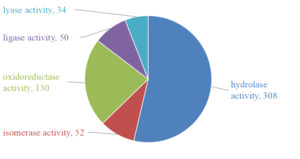
GO molecular functional analysis of the mucus proteins from Dongtingking crucian carp showed that some proteins had oxidoreductase activity, hydrolase activity, lyase activity, isomerase activity and ligase activity. The number of proteins with hydrolase activity was the most, with 308 protein sequences, and the protein with lyase activity was the least, with 34 proteins having lyase activity (Figure 3).
KEGG Analysis
KEGG analysis revealed that the skin mucus proteins were enriched in 53 KEGG pathways. These pathways included Ribosome, Bacterial infection, Phagosome, and Cell adhesion (Table 3). Specifically, the bacterial infection pathways encompassed Salmonella infection and Bacterial invasion of epithelial cells. For immune system pathways, the C-type lectin receptor signaling pathway was enriched. The Ribosome pathway involved the largest number of proteins, with 130 proteins participating, followed by the Salmonella infection pathway, which involved 107 proteins. These pathways collectively delineate an active, coordinated, multi-layered defense system. Firstly, the enrichment of the Bacterial invasion of epithelial cells pathways (involving 48 proteins, such as B-cadherin-like: A0A6P6PS88, catenin alpha-1-like:A0A6P6QRQ8, catenin beta-1-like:A0A6P6LTS4, etc.) provides molecular evidence for the reinforcement of inter-epithelial junctions beneath the mucus layer, forming the structural foundation for the physical barrier. Epithelial cells serve as a natural barrier between the external environment and internal organs, preventing entry of various external harmful factors, such as microorganisms, into the body and bloodstream. This barrier is primarily established and maintained by adherens junctions (AJs), whose main components are the cadherin/catenin “core” adhesion complex and the nectin-afadin adhesion complex.11 Together, these complexes mediate cell adhesion, maintain tissue and structural integrity, and resist microbial adhesion and invasion.12 Secondly, the enrichment of the C-type lectin receptor signaling pathway (involving 32 proteins, including calmodulin:Q6DN21, calcineurin subunit B type 1:A0A6P6JJ15, and MAPK14a:A0A6P6N9S9, etc) indicates the mucus’s capacity for immune recognition and signal transduction, enabling the initiation of conserved immune activation pathways such as CaN-NFAT in response to pathogens. Calcineurin (CaN) is the only known Ser/Thr protein phosphatase whose activity is regulated by Ca²⁺/calmodulin (CaM). The calcineurin-nuclear factor of activated T cells (CaN-NFAT) signaling pathway serves as a critical regulatory hub in T cell activation and is a key biological signaling transduction pathway in T cells.13 MAPK14 plays an important role in DNA damage and repair and can be activated by various environmental stresses and pro-inflammatory cytokines.14 Notably, the high enrichment of the Salmonella infection pathway (involving 107 proteins, including fadd:A0A6P6PT60, caspase-1-like:A0A6P6R1E4, caspase-3-like:A0A6P6N326, caspase-7-like:A0A6P6QRM7, etc.) unveils an active defense mechanism through apoptosis to eliminate infected cells, which is crucial for containing pathogen spread at the primary infection site. FADD can recruits caspase-8 to form DISC, initiating the caspase cascade, activating downstream effector caspases, and thereby triggering apoptosis to eliminate infected cells. It plays a critical regulatory role in antiviral and antibacterial immune responses.15 Furthermore, the coordinated enrichment of core metabolic pathways, namely Ribosome (130 proteins) and Glycolysis/Gluconeogenesis (40 proteins), supplies the necessary protein synthesis and acute energy required to fuel the aforementioned rapid immune responses. Glycolysis is a key metabolic pathway that not only can rapidly produce energy for immune cells, but also provides important metabolic support for maintaining their functional activities, thus promoting the activation and proliferation of immune cells.16 These enriched pathways and the involved proteins functioning in physical barrier maintenance, inflammation mediation, immune regulation, and apoptosis demonstrate that the skin mucus is a dynamic and integrated defensive interface, rather than a mere passive shield. GO annotation and KEGG pathway information for all identified mucus proteins in the proteomic study were summarized in Supplementary Table 2 via the DAVID online tool (Table S2).
Discussion
The skin serves as the first line of immune defense in fish, with mucosal immunity and skin mucus playing critical roles in survival. Fish skin mucus exhibits a complex molecular composition, and its immune functions are mediated by a diverse array of components, including antimicrobial proteins/peptides, lectins, lysozymes, and complement proteins.5 In Dongtingking crucian carp, our proteomic analysis identified numerous molecules that contribute to these protective mechanisms, which can be linked to defenses against pathogens and environmental stressors. A preliminary analysis revealed several antimicrobial proteins/peptides, such as hemoglobin subunits and histones, may belong to a class of small peptides with antimicrobial activity.17 Furthermore, various lysozymes, lectins, and complement components were detected (Table 4). These molecules establish a multi-layered antimicrobial environment: lysozymes degrade bacterial cell walls; lectins can agglutinate pathogens or enhance opsonization; and complement proteins form a cascade that can directly lyse microbes, promote phagocytosis, and trigger inflammatory responses. Notably, proteins related to substance transport and calcium-binding were abundant in the mucus of Dongtingking crucian carp. Transport-associated proteins include apolipoproteins, transferrin, hemopexin, and ceruloplasmin-like proteins, among others. Apolipoproteins, common in the skin mucus of fish,18 can lyse bacteria and exhibit strong bactericidal effects in vitro.19 Proteins such as hemopexin, transferrin, and ceruloplasmin-like proteins function as metal ion-binding proteins. By sequestering iron ions, they limit the availability of this essential nutrient for potential pathogens, employing a strategy known as nutritional immunity.20 Calcium-binding proteins identified include calmodulin, calreticulin, and annexin family proteins, among others. Calmodulin, widely distributed in fish mucus,21 is involved in maintaining osmotic balance and enhancing low-temperature resistance,22 while also participating in cellular stress and immune signaling.23 Calreticulin can bind to microorganisms and activate the phagocytic function of host phagocytes, therefore, calreticulin also plays a critical role in the immune defense of fish skin mucus.24 Notably, several of these proteins, including annexin A1-like, complement component 9, lysozyme, transferrin-a (tfa), and others participate in the biological process of response to stress. This overlap suggests that these molecules not only provide direct antimicrobial defense but also help the organism adapt to and mitigate the effects of environmental stressors, such as temperature fluctuations or pathogen challenge.
In the GO biological process analysis, the largest number of proteins (958) were enriched in metabolic processes. This is likely closely related to the maintenance of the mucus barrier function: on one hand, the continuous synthesis and renewal of key mucus components such as glycoproteins and mucopolysaccharides highly depend on active metabolic activities; on the other hand, in response to stressors like temperature and pathogens, the activation of energy metabolism pathways (e.g., TCA cycle, glycolysis) provides energy and metabolic precursors for immune responses and barrier repair. Furthermore, various metabolites (e.g., purines, organic acids) are directly involved in immune regulation. Therefore, active metabolic processes not only reflect material and energy conversion but also form the basis for mucus barrier construction, renewal, and the realization of its immune functions. Biological processes related to stress and immunity (GO:0006950 ~ response to stress) (GO:0002262 ~ myeloid cell homeostasis) were enriched with 142 and 8 mucus proteins, respectively. The former includes the previously discussed Annexin A1-like, complement component C9, lysozyme, and transferrin-a (tfa) proteins. In addition, it also includes heat shock proteins and molecular chaperones (heat shock protein 8:A0A6P6QBY3, heat shock cognate 71 kDa protein-like:A0A6P6N2I9, DnaJ heat shock protein family (Hsp40) member A2a:A0A6P6KGK7, etc.), superoxide dismutase (superoxide dismutase 1, soluble:A0A6P6Q7L9; sod3a:A0A6P6PSP1), cytoplasmic thioredoxin reductase 1-like:A0A6P6KE82, peroxiredoxin-1:A0A6P6IW66, and prothrombin-like:A0A6P6KH95, etc. In fish, heat shock proteins (HSPs) play roles in enhancing stress resistance, immune regulation, apoptosis, and inflammatory responses in aquatic organisms.25 Antioxidant enzymes such as superoxide dismutase (SOD) and peroxidase (POD) are well-recognized key enzymes in stress responses and are closely associated with the organism’s immune status. They enhance the defensive capacity of phagocytes and overall immune function, and have been identified in the skin mucus of various fish species.26–28 These proteins help mitigate oxidative damage and maintain proteostasis, thereby directly supporting the survival and function of mucus-producing cells and ensuring the integrity of the mucus barrier. Proteins enriched in the biological process of myeloid cell homeostasis include the programmed cell death protein PDCD2:A0A6P6L8H4, and Caspase family proteins such as Caspase-3-like:A0A6P6N326 and Caspase-7-like:A0A6P6QRM7, etc. PDCD2, Caspase-3, and Caspase-7 are apoptosis-related proteins. PDCD2 is believed to promote apoptosis by activating the caspase cascade reaction.29 It is thus hypothesized that in Dongtingking crucian carp, PDCD2 may help maintain myeloid cell homeostasis—including that of immune cells residing in mucus-associated tissues—by eliminating excess, aged, or damaged cells. This process likely contributes to immune surveillance and inflammation regulation, thereby indirectly preserving the stability of the mucus microenvironment.
GO Molecular function analysis revealed that some proteins possess enzymatic activity, with hydrolase activity being the most abundant among the proteins identified in Dongtingking mucus. These include lysozyme C-like (A0A6P6MZS1), lysozyme g-like isoform X2(A0A6P6QTN2), cathepsin A(ctsa) (A0A6P6PLM6), cathepsin D(A0A6P6RBJ5), chymotrypsin-like protease CTRL-1(A0A6P6PF40), among others. Studies have reported hydrolases in fish skin mucus are one of the primary substances responsible for nonspecific immunity.30 Hydrolases in the mucus can dissolve parasites on the fish body surface, thereby protecting the organism.5 In fish mucus, the composition of protein is very complex. Mucins such as mucin-2-like:A0A6P6MDT6 and mucin-5AC-like:A0A6P6PKF2 have been identified in Dongtingking crucian carp mucus. Mucin, as a component of mucus, plays a significant role in fish’s resistance to pathogen invasion. They carry numerous oligosaccharide chains that undergo molecular cross-linking and bind water molecules, forming a slippery surface layer on the fish skin.4 Additionally, other potentially immune-related proteins have been found in Dongtingking crucian carp mucus, including Vitellogenin-like:A0A6P6J7E6, alpha-2-macroglobulin-like:A0A6P6QYB1, etc.
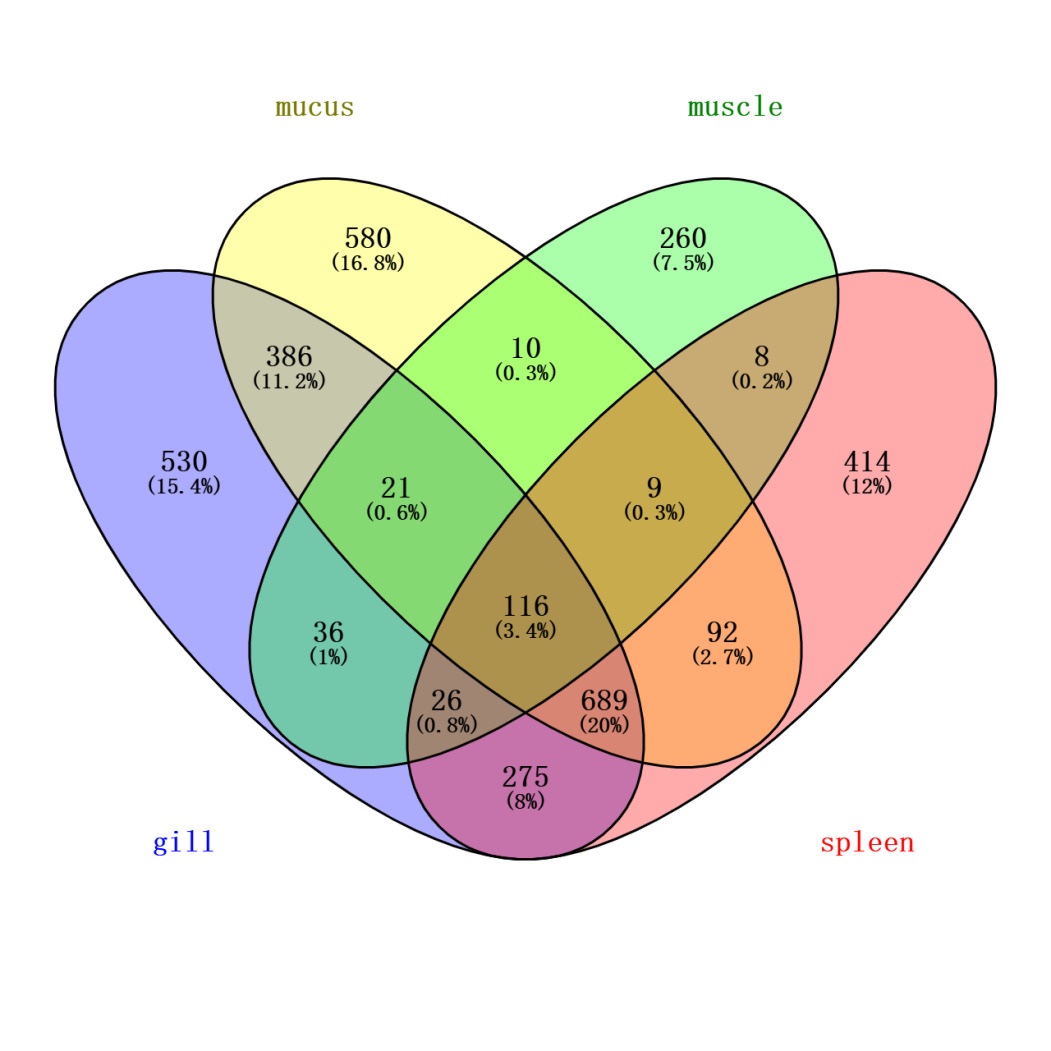
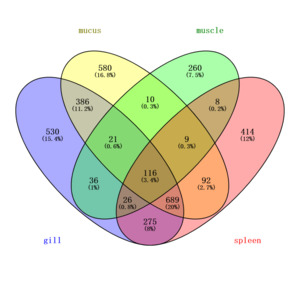
To more accurately identify mucus-specific protein components, the skin mucus protein samples of Dongtingking crucian carp were processed and sequenced alongside samples from gill, spleen, and muscle tissues. Venny 2.1 software was employed to identify proteins unique to these four types of samples (Figure 4). The analysis revealed 580 proteins exclusive to the mucus samples, including Mucin-5AC-like:A0A6P6PKF2, the lectin Malectin:A0A6P6MTU9, the calcium sensor protein Peflin:A0A6P6NWU5, the cell junction protein Pinin-like:A0A6P6NJS8, T-cell surface antigen CD2-like:A0A6P6NIU6, sex hormone-binding globulin-like:A0A6P6QHK5, translin-like:A0A6P6Q119, and FAS-associated death domain protein (FADD):A0A6P6PT60, among others. We recognize that while the synchronous processing and sequencing of multiple tissue samples facilitates experimental consistency, it carries a theoretical risk of cross-contamination. To minimize this risk, targeted measures were implemented during sample collection. Specifically, during the sampling process, mucus was collected with utmost care to avoid contamination from gill mucus or excretions. Following mucus collection, other tissues were separately harvested using sterile scissors and forceps. The list of “mucus-specific proteins” obtained through Venny analysis of proteins identified across all tissues, represents a key analytical step taken to exclude, as thoroughly as possible, common proteins including potential contaminants. It is worth noting that this list includes recognized key structural components of mucus, such as Mucin-5AC-like, as well as a series of proteins functionally implicated in mucosal immunity, barrier defense, and environmental sensing (e.g., the aforementioned lectins and calcium-sensing proteins). The specific enrichment of these proteins in the mucus samples, coupled with the high congruence of their biological functions with the theoretical roles of mucus, provides strong corroborating evidence for the specificity of our data from a biological standpoint.
The complex protein composition in the mucus enables fish to efficiently counteract the ubiquitous pathogens and physicochemical challenges in the aquatic environment. Proteomics technology was used to search for immune-related proteins in Dongtingking crucian carp mucus, which had the advantages of wide coverage and high accuracy. Although these immune-related proteins in the mucus have been successfully identified, further in-depth research is necessary to explore their correlations with piscine diseases, thereby advancing the prevention and treatment of fish diseases.
The protein composition in fish mucus is highly complex. In this experiment, chromatography-mass spectrometry techniques were employed to separate and identify proteins in the mucus of Dongtingking crucian carp, and a total of 1,903 mucus proteins were identified. Among these, antimicrobial proteins/peptides, lectins, lysozymes, complement components, transport-related proteins, calcium-binding proteins, and superoxide dismutase were identified in the skin mucus of Dongtingking crucian carp. GO enrichment analysis revealed that the skin mucus proteins of Dongtingking crucian carp were primarily enriched in biological processes such as metabolic processes, cellular component organization or biogenesis, macromolecule localization, cellular localization, and response to stress. KEGG analysis showed that these mucus proteins are mainly involved in pathways including ribosome, bacterial infection, phagosome, and cell adhesion. Based on the proteomic data of Dongtingking crucian carp, further work will focus on functional studies of mucus proteins and protein-protein interactions. We will prioritize experimental validation of the bacteriolytic activity of proteins with direct antimicrobial potential—such as the identified complement components, lysozymes, and antimicrobial peptides—against major aquatic pathogens. Furthermore, we will emphasize investigating the regulatory functions of mucus-specific proteins in bacterial infection and immune-related pathways, including Salmonella infection, bacterial invasion of epithelial cells, and the C-type lectin receptor signaling pathway. These studies aim to elucidate the mechanisms of action of the relevant proteins, thereby providing a theoretical foundation for improving the prevention of fish diseases.
Acknowledgments
We thank Ms. Min Wang from the Sequencing Platform of the Institute of Hydrobiology, Chinese Academy of Sciences, for her assistance in the mass spectrometric analysis.
The research was supported by the Excellent Young Scholars Program of the Hunan Provincial Education Department (No. 23B0662), the General Project of Hunan Provincial Natural Science Foundation (No. 2025JJ50144), and the Scientific Research Project of Hunan University of Arts and Science (No. 24YB06). Acknowledgement is made to the above funds for their support.
Authors’ Contribution
Conceptualization: Bolan Zhou (Equal), Pinhong Yang (Equal), Wei Peng (Equal). Methodology: Bolan Zhou (Equal), Haoren Yang (Equal), Pinhong Yang (Equal), Yunsheng Zhang (Equal). Investigation: Bolan Zhou (Equal), Haoren Yang (Equal), Yunsheng Zhang (Equal). Software: Bolan Zhou (Equal), Haoren Yang (Equal), Xianyuan Zhou (Equal). Writing – original draft: Bolan Zhou (Equal), Xianyuan Zhou (Equal). Funding acquisition: Bolan Zhou (Equal), Yunsheng Zhang (Equal), Wei Peng (Equal). Writing – review & editing: Bolan Zhou (Equal), Pinhong Yang (Equal), Wei Peng (Equal), Xianyuan Zhou (Equal). Supervision: Bolan Zhou (Equal), Wei Peng (Equal), Xianyuan Zhou (Equal). Data curation: Haoren Yang (Lead). Resources: Pinhong Yang (Lead).
Competing Interests
No competing interests were disclosed.
Ethical Conduct Approval – IACUC
All animal experiments were conducted in accordance with the Guiding Principles for the Care and Use of Laboratory Animals and were approved by Hunan University of Arts and Science.
Informed Consent Statement
All authors and institutions have confirmed this manuscript for publication.
Data Availability Statement
The mass spectrometry proteomics data generated in this study have been deposited to the iProX integrated proteomics resource (accession: IPX0014666000) and the ProteomeXchange Consortium (accession: PXD071967). The data are currently private and the dataset will be made publicly available upon publication of the associated article.

_enrichment_analysis_of_skin_mucus_proteins_in_dongtingking_crucian_carp.tiff)
_enrichment_analysis_of_skin_mucus_proteins_in_dongtingking_crucian_carp.tiff)