INTRODUCTION
In China, the crayfish (Procambarus clarkii) has gained significant economic importance, with its farming area expanding to 1.97 million hectares and production reaching 3.16 million tons by 2023.1 However, this rapid growth, combined with recurring water quality issues, has led to frequent outbreaks of viral and bacterial diseases.2 Among these, white spot syndrome is regarded as one of the most destructive viral diseases affecting crustaceans, including crayfish.3 White spot syndrome virus (WSSV), the etiological agent of the disease, is a double-stranded DNA virus classified under the family Nimaviridae and the genus Whispovirus. The virions are rod-shaped, measuring approximately 250–380 nm in length and 80–120 nm in width, and possess a flagellum-like appendage. It is now understood that WSSV is highly virulent and rapidly transmissible, capable of infecting a wide range of decapod crustaceans, including white leg shrimp (Litopenaeus vannamei), crayfish, and various crab species. The disease also causes high mortality rates and significant economic losses for the global aquaculture sector.4 Currently, there are no approved drugs or treatment strategies for WSSV, highlighting the urgent need for effective preventive or therapeutic solutions.
Plants offer a valuable source of low-cost, safe phytochemicals with a broad range of pharmacological properties, including antiviral effects.5 Growing attention has been directed toward harnessing the plants or plant-sourced compounds for the prevention and control of WSSV in crustacean farming. Recent research has shown that different methanol extracts and active constituents from plants such as rhein, geniposidic acid, paeonol, and Japanese honeysuckle (Lonicera japonica) exhibit potent anti-WSSV effects in crayfish.6–9 These findings underscore the practicality and efficiency of identifying plant-derived compounds with antiviral properties as a low-cost strategy for discovering promising candidates against WSSV.
Scutellaria baicalensis (SB), a perennial medicinal plant from the Lamiaceae family, has demonstrated multiple pharmacological benefits in managing diseases in aquatic animals, including fish and crustaceans.10 Flavonoid compounds from this plant have been reported to harbor notable antibacterial effects (e.g., against Aeromonas hydrophila) and antiviral properties (including activity against WSSV) and can also enhance the health status of aquatic animals through antioxidant and immunomodulatory mechanisms.11,12 Flos Sophorae Immaturus (FSI), the dried blossom bud of Flos Sophorae Immaturus L., contains an array of bioactive compounds with multiple pharmacological benefits, including antithrombotic, anticancer, analgesic, antimicrobial, antiviral, anti-aging, anti-inflammatory, antioxidant, antiallergic, and hypoglycemic effects.13 Prunella vulgaris (PV), a fragrant perennial herb, reportedly possesses immunomodulatory, antioxidant, antiviral, anti-inflammatory, anti-allergic, and anticancer properties, with the most potent antiviral effect observed against fish viruses like infectious hematopoietic necrosis virus (IHNV).14,15 Herbal compounds demonstrate unique potential in the antiviral field due to their multi-component and multi-target synergistic effects.16 The SB, FSI, and PV used in this study all possess promising pharmacological properties. Although some studies have revealed the inhibitory effects of these individual herbs against pathogens such as WSSV and IHNV, comprehensive research on the antiviral effects of their compound formulations and their synergistic mechanisms in combined form remains scarce. Given their diverse mechanisms, encompassing antioxidant, immunomodulatory, and antimicrobial activities, exploring their combined use may offer a novel strategy to enhance viral resistance in aquatic species. Further research is warranted to comprehensively evaluate their potential for antiviral therapy in aquaculture.
In the present study, crayfish were employed as an experimental model to evaluate the antiviral effects of a compound herbal formula comprising SB, FSI, and PV against WSSV. Our investigation focused on viral load, histopathological alterations, and the expression of innate immune-related genes in the hepatopancreas, intestine, and gills. Furthermore, changes in intestinal microbiota composition and metabolomic profiles were assessed. Overall, the results of this study offer potential candidate therapeutics for WSSV management in aquaculture and contribute to the scientific advancement of traditional Chinese medicine-based polyherbal formulations.
MATERIALS AND METHODS
CRAYFISH AND HUSBANDRY PROTOCOL
Crayfish (mean weight 20 ± 1.03 g) were obtained from a fish farm in Wuhan, Hubei Province, China. Before experimentation, the animals were screened for WSSV via conventional PCR and confirmed to be WSSV-free. Following a 14-day acclimation period, the crayfish were randomly allocated into four experimental groups (n = 50 per group): control (CON), virus challenge (CHG), drug-treated (DTG), and treatment (TRE) groups. The DTG group was included to control any potential toxic effects of the herbal compound (in comparison with the CON group) and to help distinguish its specific immunostimulatory or antiviral effects from non-specific physiological impacts (in comparison with the TRE group). The CON and CHG groups administered a standard diet, while the DTG and TRE groups received a medicated diet containing the herbal compound. After 7 days of feeding, crayfish in the CHG and TRE groups received intramuscularly injections of 100 μL of WSSV suspension. All groups were then maintained for an additional 7 days, for a total duration of 14 days (Figure 1A). During the rearing period, feeding was conducted twice daily at 09:00 and 17:00. Partial water exchange (50% of the total volume) was performed every 48 hours. Daily observations were performed to monitor survival and feeding behavior. Environmental stability was maintained using shade nets, heating rods, and continuous aeration systems. All animal experiments were conducted in compliance with the guidelines of the Animal Ethics Committee of Yangtze River Fisheries Research Institute, Chinese Academy of Fishery Sciences, and the protocol was approved under the ethical permission code: YFI2024-zhouyong-11
HERBAL COMPOUND PREPARATION AND WSSV CHALLENGE
The herbal compound, consisting of SB, FSI, and PV at a weight ratio of 2:2:1 (optimized through preliminary in vitro antiviral screening), was subjected to water extraction. The aqueous extract was concentrated, spray-dried to a powder, and standardized to a total flavonoid content of ≥15% (w/w). This standardized powder was incorporated into the basal feed at 1:100 (w/w) to prepare the medicated diet. The WSSV strain used for viral challenge was previously isolated and characterized by our laboratory.
TISSUE SAMPLING AND PRESERVATION
Feeding was ceased 24 hours before sampling. From each experimental group, six crayfish were randomly selected, and their hepatopancreas, intestine, and gill tissues were dissected and promptly conserved -80 ℃ for subsequent DNA and RNA extraction. An additional six individuals per group were sampled for histological analysis. The same tissues (hepatopancreas, intestine, and gills) were excised, rinsed thoroughly with PBS to remove residual blood, and fixed in appropriate fixatives for Alcian Blue (AB) and Hematoxylin and Eosin (HE) staining. Furthermore, six additional crayfish were collected per group for gut microbiota and metabolomic analyses. Under sterile conditions, intestinal tissues were dissected, rinsed using sterile distilled water, gently blotted dry with sterile filter paper to remove surface moisture, snap-frozen in liquid nitrogen, and conserved at -80℃ until subsequent processing
QUANTIFICATION OF WSSV LOAD IN CRAYFISH TISSUES
Quantitative PCR (qPCR) with a standardized plasmid was used to determine WSSV copy numbers. The plasmid contained a conserved region of the WSSV genome for accurate quantification. A standard curve was generated based on consecutive dilutions of the plasmid with known copy numbers and their corresponding cycle threshold (Ct) values, from which a linear regression equation was established. Primer sequences are provided in Table 1. Genomic DNA was isolated from tissue samples following the Yeasen Kit protocol. DNA concentration and purity were measured with a micro-volume spectrophotometer, and all samples were subsequently normalized to 200 ng/μL. The absolute viral load was quantified using qPCR and the standard curve.
PREPARATION OF HISTOPATHOLOGICAL SECTIONS
Hepatopancreas, intestinal, and gill tissues were collected, fixed, embedded in paraffin, and sectioned using a microtome. For HE staining, tissue sections were deparaffinized and rehydrated. They were then stained with hematoxylin for nuclear visualization, followed by a differentiation step and bluing. The sections were subsequently stained with eosin to highlight the cytoplasm and extracellular matrix. After staining, the sections were dehydrated, cleared, and mounted with coverslips for microscopic observation. For AB staining, intestinal and gill sections were similarly processed. Following fixation, paraffin embedding, sectioning, deparaffinization, and rehydration, sections were stained with Alcian Blue, rinsed, counterstained with nuclear fast red, dehydrated, cleared, and cover-slipped. All treated sections were subsequently examined under a light microscope to evaluate histopathological alterations.
IMMUNE GENE EXPRESSION PROFILING
Total RNA was isolated from tissue samples using the Yeasen Cell/Tissue RNA Extraction Kit, adhering to the manufacturer’s guidelines. The RNA concentration and purity were measured with a micro-volume spectrophotometer. Subsequently, cDNA was synthesized from the extracted RNA using the Yeasen Reverse Transcription Kit based on the provided instructions. RT-qPCR was conducted using the synthesized cDNA as a template. The primer sequences utilized in this study are detailed in Table 1. Relative gene expression levels were computed employing the 2-ΔΔCt method.
GUT MICROBIOTA ANALYSIS
The intestinal microbiota composition was evaluated using 16S rRNA gene sequencing. Total bacterial DNA was isolated and purified from gut samples utilizing the OMEGA Soil DNA Kit (D5635-02; Omega Bio-Tek, Norcross, GA, USA), adhering to the manufacturer’s guidelines. The V3–V4 hypervariable regions of the 16S rRNA gene were PCR-amplified with primers 341F (5’-GTACTCCTACGGGAGGCAGCA-3’) and 806R (5’-GTGGACTACHVGGGTWTCTAAT-3’). PCR conditions were as follows: initial denaturation at 98 ℃ for 5 min; 25 cycles of 98 ℃ for 30 s, 53 ℃ for 30 s, and 72 ℃ for 45 s; followed by a final extension at 72 ℃ for 5 min. Amplification products were stored at 12 ℃ until further processing. PCR amplicons were resolved on a 2% agarose gel, and the target bands were excised and purified using the Axygen Gel Extraction Kit. The purified amplicons were quantified using the Quant-iT PicoGreen dsDNA Assay Kit on a BioTek FLx800 microplate reader. Equal molar amounts of amplicons were pooled to achieve the desired sequencing depth. Sequencing libraries were prepared with the Illumina TruSeq Nano DNA LT Library Prep Kit and underwent size selection and purification on a 2% agarose gel. Concentrations were quantified using the Qubit fluorometer and verified by quantitative PCR. Following quality control, the libraries were sequenced on an Illumina NovaSeq platform. Raw sequencing data were filtered and denoised before clustering into operational taxonomic units (OTUs). Taxonomic assignments were performed with standard reference databases. Microbial community composition was quantified and compared across samples at various taxonomic levels. Data normalization and calculation of alpha and beta diversity indices were conducted using R software and QIIME2 pipelines.
GUT METABOLOMICS ANALYSIS
Untargeted metabolomic profiling was conducted using a Thermo Scientific high-definition liquid chromatography-mass spectrometry (LC-MS) platform. Intestinal tissues and luminal contents from crayfish were homogenized in 80% methanol (v/v, pre-chilled to -20℃) at a tissue-to-solvent ratio of 1:10 (w/v), centrifuged at 15,000 × g for 15 min at 4°C to obtain metabolite extracts for LC-MS analysis. Metabolite identification and quantification were performed using Compound Discoverer™ 3.3 software (Thermo Fisher Scientific), in conjunction with multiple MS/MS spectral libraries (mzCloud, ChemSpider, and LipidBlast).
Chromatographic separation was performed on an ACQUITY UPLC HSS T3 column (100 Å pore size, 1.8 μm particle size, 2.1 × 100 mm; Waters) under the following conditions: mobile phase flow rate of 0.4 mL/min, column temperature maintained at 40℃, autosampler temperature at 8℃, and injection volume of 2 μL. The mobile phase consisted of 0.1% (v/v) formic acid in water (solvent A, positive ion mode) and 0.1% (v/v) formic acid in acetonitrile (solvent B, negative ion mode). The detailed gradient elution program is provided in Table 2.
Mass spectrometric analysis was performed using a Thermo Orbitrap Exploris 120 mass spectrometer driven by Xcalibur 4.7 software, operating in data-dependent acquisition (DDA) mode for both positive and negative ionization. The heated electrospray ionization (HESI) source was operated with the following parameters: spray voltage of ±3.5 kV (positive mode)/-3.0 kV (negative mode), sheath gas flow of 40 arbitrary units, auxiliary gas flow of 15 arbitrary units, capillary temperature of 325°C, and temperature of the auxiliary gas heater of 300°C. Full scan MS spectra were acquired at a resolution of 60,000 (scan range m/z 100-1000) with an automatic gain control (AGC) target of “Standard” and maximum injection time of 100 ms. For MS/MS analysis, the four most intense precursor ions per cycle were selected for higher-energy collisional dissociation (HCD) fragmentation (normalized collision energy: 30%) at a resolution of 15,000, with dynamic exclusion set to 8 s, the AGC target to “Standard”, and the maximum injection time was automatically determined.
INTEGRATED ANALYSIS OF GUT METABOLOMICS AND MICROBIOTA DATA
The R package ropls was used to conduct multivariate statistical analyses such as principal component analysis (PCA), partial least squares discriminant analysis (PLS-DA), and orthogonal partial least squares discriminant analysis (OPLS-DA) to determine the variable importance in projection (VIP) scores for each metabolite. Metabolites with p < 0.05 and VIP > 1 were considered statistically significant. The abundance values of the identified differential metabolites were subjected to hierarchical clustering analysis using the pheatmap package (v1.0.12) in R. KEGG pathway enrichment analysis of differential metabolites was performed using cluster Profiler (v4.6.0) to identify significantly enriched metabolic pathways. Venn diagrams and Upset plots for pairwise comparisons were generated using Venn Diagram (v1.7.3) and UpSetR (v1.4.0), respectively. Furthermore, to delineate the co-regulation patterns established after treatment, a Spearman correlation analysis was performed specifically within the TRE group, focusing on the beneficial metabolites and microbial genera (including both up-regulated beneficial and down-regulated harmful genera) that were significantly altered compared to the CHG group.
RESULTS
THE COMPOUND FORMULATION INHIBITS WSSV REPLICATION
At 7 days post-challenge, the survival rate of crayfish in the CHG group was significantly lower than in the TRE group, demonstrating that the compound formulation composed of SB, FSI, and PV significantly reduced WSSV-induced mortality. Furthermore, throughout the experimental period, the survival rate of the DTG group showed no significant difference compared to the CON group. Daily behavioral observations further confirmed normal feeding activity and swimming behavior in both the DTG and CON groups (Figure 1B). A standard curve was established using a WSSV plasmid with known copy numbers, yielding the equation y = -0.2395x + 10.66 (R² = 0.9943), indicating excellent linearity (Figure 1C). qPCR analysis at 7 days post-challenge revealed that WSSV viral loads in hepatopancreas, intestinal, and gill tissues of the TRE group were significantly lower than those in the CHG group, validating the compound’s efficacy in inhibiting WSSV replication in crayfish (Figure 1D-F).
THE EFFECTS OF COMPOUND MEDICINE ON THE HISTOLOGICAL STRUCTURE OF CRAYFISH TISSUES
To further substantiate the antiviral efficacy of the compound medicine composed of SB, FSI, and PV against WSSV, we performed HE staining and histopathological observations on the hepatopancreas, intestine, and gill tissues of crayfish.
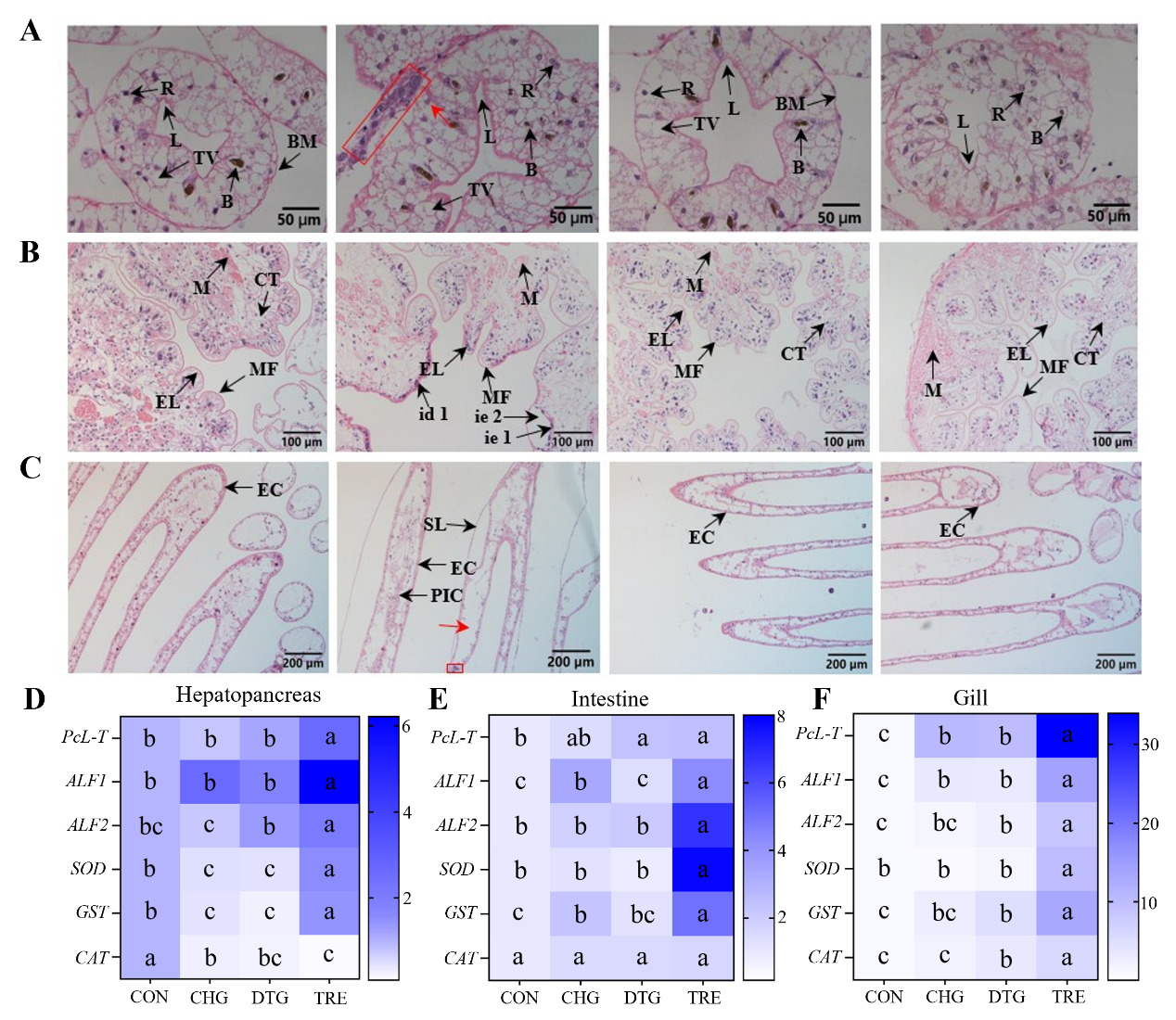
In the control group (CON), hepatopancreatic cells exhibited normal histological architecture, characterized by uniform cell distribution, normal volume of blasenzenllen cells (B cells) and their intracellular transport vesicles, intact hepatic tubules, and no signs of pathological changes such as inflammatory infiltration or cell necrosis. The virus-challenged group (CHG) exhibited significant pathological alterations, including dilated hepatic tubule lumens, extensive inflammatory cell infiltration (indicated by boxed area), enlarged storage cells, focal necrosis (indicated by red arrow), and reduced volume of both transport vesicles and B cells. Notably, the treatment group (TRE) showed significant tissue repair, characterized by reduced storage cell volume, a return of most storage cells to a morphology resembling the control state, restoration of hepatic tubule structure, absence of inflammatory or necrotic cells, and normal B cell size (Figure 2A).
In addition, in the CON group, intestinal villi were formed by the invagination of epithelial cells and the submucosa, arranged neatly, with intact gill filament structure, with no hemocoel dilation or inflammatory cell infiltration, while the secondary lamellae exhibited signs of atrophy. In contrast, the CHG group exhibited detachment and apparent damage of epithelial tissue from the connective tissue, along with atrophy of intestinal epithelial cells and shriveled, disordered villi. Moreover, the hemocoels of the gill filaments were severely dilated (indicated by red arrows), with many inflammatory cells present in the hemocoel (boxed area). The columnar cells were disorganized, and the secondary lamellae were hyperplastic. Conversely, in the TRE group, intestinal epithelial cell atrophy was absent, intestinal villi arrangement became more orderly, and damage to the epithelial layer was resolved. Moreover, the hemocoel of the gill filaments returned to a state similar to the CON group, inflammatory cells were absent, the secondary lamellae were atrophied, and the columnar cells were arranged in an orderly manner (Figure 2B-C). These results validated that the compound formulation composed of SB, FSI, and PV could effectively alleviate histopathological damage to the hepatopancreas, intestine, and gills caused by WSSV, significantly mitigating inflammatory responses, inhibiting cell necrosis, and promoting tissue repair. Meanwhile, AB-stained pathological sections showed that, relative to the CHG group, mucus secretion in the intestinal and gill tissues of crayfish in the TRE group was significantly reduced, further corroborating this finding (Appendix Figure 1A-B). The tissue morphology of the DTG group exhibited no significant difference from that of the CON group, indicating the low toxicity of this compound formulation to crayfish.
THE IMPACT OF COMPOUND MEDICATIONS ON THE IMMUNE GENES OF CRAYFISH
As crustaceans like the red swamp crayfish rely solely on innate immunity for antiviral defense, we investigated whether the SB-FSI-PV compound could enhance this crucial immune arm. RT-qPCR analysis revealed significant upregulation of five innate immune genes (PcL-T, ALF1, ALF2, SOD, and GST) in the hepatopancreas, intestinal, and gill tissues of the TRE group compared to the other three groups (CON, CHG, and DTG) in crayfish (p < 0.05). A similar expression pattern was observed for the CAT gene in both intestinal and gill tissues (p < 0.05). These findings suggest that the compound formulation composed of SB, FSI, and PV may exert its antiviral effects through the modulation of these immune-related genes (Figure 2D-F).
MICROBIAL DIVERSITY AND COMMUNITY STRUCTURE PROFILING OF THE GUT MICROBIOME IN CRAYFISH
Alpha diversity analysis serves as a crucial indicator for assessing intestinal microbial community diversity in crayfish, primarily focusing on species richness and evenness within individual samples. The Chao1 index and Shannon index represent species richness and evenness, respectively. Comparative analysis of these indices across treatment groups revealed the highest values in the CHG group, followed by TRE, DTG, and CON groups, demonstrating that both WSSV infection and drug treatment significantly affected the diversity and uniformity of the intestinal microbiota in crayfish. (Figure 3A-B).
Beta diversity analysis represents a fundamental approach for examining differences in microbial community composition between samples. Principal Coordinates Analysis (PCoA), a widely used dimension-reducing method, reflects similarities or differences in community composition among samples. Smaller inter-sample distances indicate greater similarity in bacterial community structure, while point-to-point distances reflect the degree of dissimilarity. Notably, sample points within each group (except CHG) showed relatively dispersed distributions, suggesting variations in microbial community structure among the three replicates. While the sample points connections for the CON, DTG, and TRE groups exhibited some overlap, no such overlap occurred with the CHG group, indicating substantial differences in the composition of microbial communities between CHG and the other three groups (Figure 3C).
At the phylum level, the top 5 most abundant phyla across all groups primarily included Proteobacteria, Firmicutes, Bacteroidota, Actinobacteriota, and Patescibacteria. In the CHG group, Proteobacteria exhibited the highest abundance, approximately twice that of the CON group, followed by the TRE group, with the DTG group showing the lowest abundance. Firmicutes demonstrated the highest abundance in the DTG group, about three times that of the TRE group and six times that of the CHG group, with the CON group having a slightly lower abundance than the DTG group. The abundance ratio of Bacteroidota among the four groups was CON: CHG: DTG: TRE = 19:33:32:24, while the abundance of Actinobacteriota and Patescibacteria was relatively low across all four groups, each being below 5%. Among them, the CON group exhibited a markedly higher abundance than the other three groups (Figure 3D).
At the genus level, the only dominant genus (abundance >5%, same below) common to all groups was Bacteroides-H, with the largest proportion in the CHG group (22.22%), followed by the CON group (17.19%), the DTG group (20.11%), and the TRE group (19.79%). Citrobacter-A was a shared dominant genus among the CON, CHG, and TRE groups, with corresponding proportions of 16.7%, 11.45%, and 32.57%, respectively, but was present at a significantly lower proportion in the DTG group (2.2%). Streptococcus and Anaerorhabdus were the dominant genera in the CON (22.83%) and DTG (8.34%) groups, respectively. Sphingobacterium was the predominant genus in both the CHG (8.63%) and DTG (5.34%) groups. Shewanella was the dominant genus in the CON group (5.52%). Besides, the CHG group did not exhibit the presence of Streptococcus, Propionivibrio, while the CON group lacked Propionivibrio. Dysgonomonas, Kaistella, and Pseudomonas-E were common across all groups, although their relative abundance remained below 5%. Specifically, the proportions of Dysgonomonas and Kaistella in the DTG group were relatively high (2.76% and 2.92%, respectively), while Pseudomonas-E was most abundant in the CHG group (3.37%) (Figure 3E).
It is well-established that the count of unique OTUs reflects the complexity of the gut microbiome. In the present study, the CON, CHG, DTG, and TRE groups exhibited 462, 1,407, 1,206, and 1,365 unique OTUs, respectively. Compared to the other three groups, the CON group exhibited substantially fewer unique OTUs, indicating a simpler gut microbial community structure and confirming that WSSV infection and medication use impact the richness of the gut microbiome in crayfish (Figure 3F).
THE COMPOUND FORMULATION PROMOTES INTESTINAL ECOLOGICAL BALANCE IN CRAYFISH
To further investigate the effects of WSSV infection and drug treatment on crayfish gut health, we generated a heatmap comparing the top 30 most abundant bacterial genera, visualizing the abundance variations of each genus across different groups (Figure 4A). At the phylum level, relative to the CON group, the Firmicutes/Bacteroidetes (F/B) ratio decreased in all three other groups, with the CHG group experiencing the most significant reduction. This indicated that WSSV infection could affect the F/B ratio, while the compound medicine mitigated this impact (Figure 4B). Besides, the heatmap revealed that compared to the CHG group, potentially harmful genera such as Bacteroides-H, Shewanella, and Pseudomonas-E were reduced in the TRE group, while beneficial genera like Anaerorhabdus and Propionivibrio were increased (Figure 4C-G). Taken together, these findings demonstrate that the compound medicine composed of SB, FSI, and PV improves crayfish intestinal health.
ANALYSIS OF DIFFERENCES IN INTESTINAL METABOLITES OF CRAYFISH
PCA revealed that PC1 and PC2 contributed 20.5% and16.9% of the variance, respectively, for a cumulative contribution of 37.4%. The samples within the CON group were more dispersed, followed by the CHG group. The samples within the DTG and TRE groups were more clustered, indicating that the effects of WSSV infection and medication resulted in a reduction in metabolic differences among samples within each group. Moreover, the overlap observed among the samples of the four groups suggests distinct metabolic patterns between them (Figure 5A).
Using the PLS-DA model, the distribution patterns within each group were consistent with the PCA outcomes. However, the overlap between the CON and CHG groups was no longer observed, indicating that WSSV infection increased their metabolic differences. The observed overlap between the DTG and TRE groups highlights the drug’s ability to restore metabolic differences between groups induced by WSSV infection (Figure 5B).
OPLS-DA, a form of multivariate statistical analysis, was employed in the present study to highlight distinctions among groups within the model, where the proximity of data points indicates similarity in metabolic profiles. OPLS-DA was employed to independently examine disparities between the CON and DTG groups, as well as between the CHG and TRE groups. The analysis revealed substantial differences, indicating that the treatment modified the gut metabolite composition in crayfish, likely related to alterations in the richness of the gut microbiome triggered by the medication (Figure 5C-D).
IDENTIFICATION AND SCREENING OF DIFFERENTIAL METABOLITES IN THE GUT OF CRAYFISH
The classification of gut metabolites across groups revealed that lipids and lipid-like molecules comprised the largest proportion (28.7%), followed by organic acids and derivatives (26.4%), organoheterocyclic compounds (15.3%), organic oxygen compounds (6.9%), phenylpropanoids and polyketides (2.3%), alkaloids and derivatives (1.4%), benzenoids (10.5%), organic nitrogen compounds (4.7%), nucleosides, nucleotides and analogs (2.2%), and organic sulfur compounds (0.62%) (Figure 6A).
Besides, significant differences were observed in the types of gut metabolites among the groups. A total of 85 metabolites were differentially expressed between CON and CHG groups, with 38 upregulated and 47 downregulated metabolites. Similarly, 149 metabolites were differentially expressed between CON and DTG groups, with 65 upregulated and 84 downregulated metabolites. 75 metabolites showed differential expression between DTG and TRE groups, with 19 upregulated and 56 downregulated. 105 metabolites were differentially expressed (26 upregulated and 79 downregulated) between CHG vs. TRE groups. The changes in metabolites suggest that under the influence of WSSV infection and medication, harmful metabolites and antiviral-related metabolites were produced (Figure 6B). A petal map of differential metabolites was generated to visualize the number of shared metabolites among groups, with the fewest shared metabolites between DTG and TRE (n = 14), and the most shared metabolites between CON and TRE (n = 47), indicating that the compound medicine composed of SB, FSI, and PV could effectively reverse the metabolic disorder caused by WSSV infection, restoring gut metabolite composition towards a healthier state (Figure 6C).
By conducting a differential analysis of metabolites between the CHG group and the TRE group, the primary biochemical metabolic pathways and signaling pathways involved were identified. The darker the red hue, the higher the likelihood of the pathway’s overall expression being upregulated; in contrast, the darker the blue hue, the higher the likelihood of the pathway’s overall expression being downregulated. The downregulated pathways mainly included cholesterol metabolism, bile secretion, cortisol synthesis and secretion, digestion and absorption of fats, aldosterone metabolism, and vitamin metabolism. The upregulated pathways included ether lipid metabolism, thiamine metabolism, beta-alanine metabolism, glycerophospholipid metabolism, pyrimidine metabolism, arachidonic acid metabolism, and the biosynthesis of pantothenate, CoA, insect hormones, primary bile acids, steroids, and steroid hormones (Figure 6D).
TARGETED ASSOCIATION BETWEEN GUT MICROBIOTA AND METABOLITES
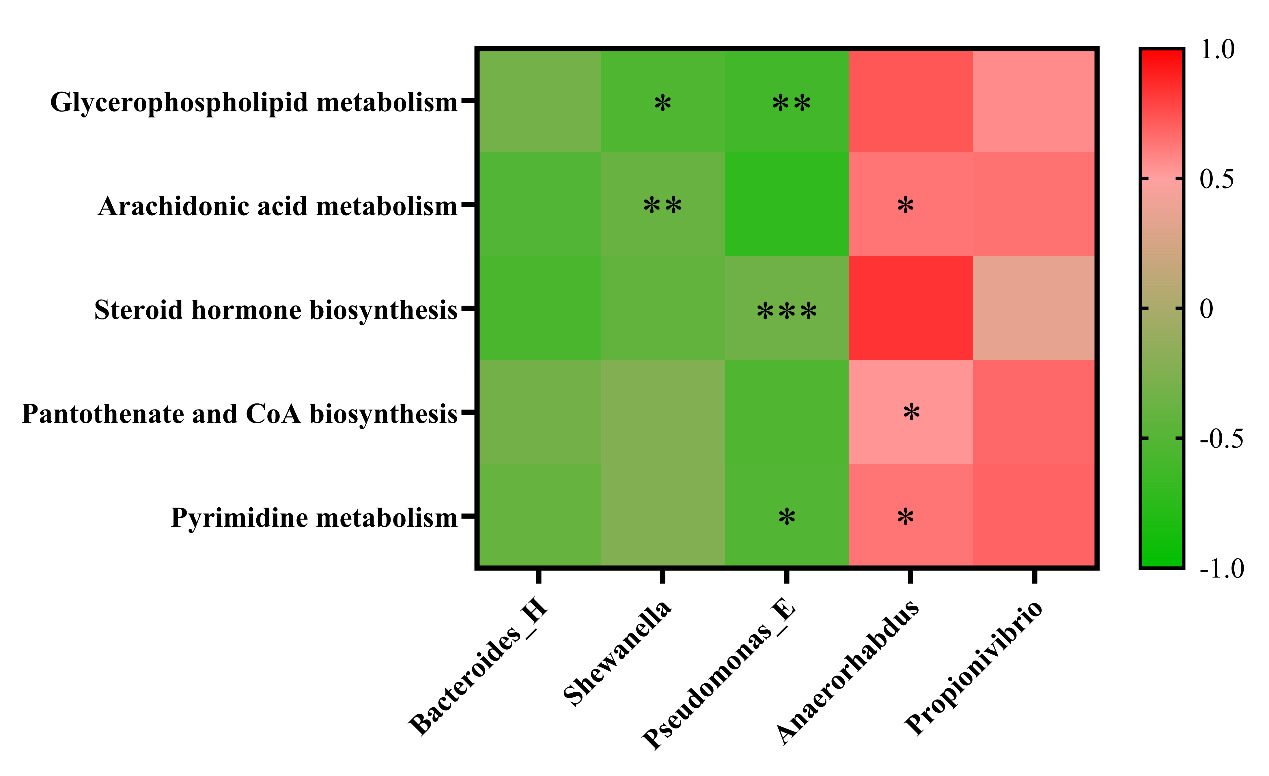
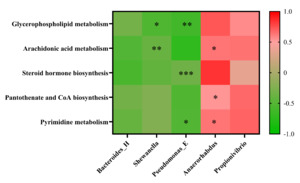
To elucidate the coordinated regulatory network established following the intervention with the herbal compound, we performed Spearman correlation analysis using experimental data from the TRE group. We observed a significant and consistent association pattern: herb-enhanced beneficial bacteria (such as Anaerorhabdus and Propionivibrio) showed significant positive correlations with multiple upregulated beneficial metabolites (including compounds from glycerophospholipid metabolism, arachidonic acid metabolism pathways, and CoA biosynthesis precursors) (|ρ| > 0.6, p < 0.05). Conversely, harmful bacteria that proliferated aberrantly during WSSV infection (such as Shewanella and Bacteroides-H) exhibited negative correlation trends with these beneficial metabolites (Appendix Figure 2).
DISCUSSION
In contrast to synthetic chemicals, herbal preparations are environmentally friendly and leave no harmful residues, positioning them as attractive alternatives for sustainable disease management in aquaculture.17,18 While much of the current research has focused on the pharmacological effects of individual herbal compounds or single plant extracts, these may exhibit limited ability to address complex aquatic diseases caused by multifactorial challenges such as viral-bacterial coinfections and environmental stressors. In such scenarios, multi-herb formulations with complementary and synergistic actions hold greater promise. However, systematic investigations into the use of compound herbal remedies for aquatic animal health remain scarce globally. In this study, a novel formulation combining SB, FSI, and PV was developed, drawing on their diverse antiviral properties. Specifically, SB polysaccharides are known to enhance immune responses, quercetin derived from FSI exhibits potent inhibition of viral enzymes, and PV extracts exert protective effects by modulating inflammatory pathways.19–21 This rational combination aims to achieve synergistic efficacy, offering a scientific foundation for the future development of compound herbal products in aquaculture.
Our findings demonstrated that a compound formulation comprising SB, FSI, and PV could reduce the viral load of WSSV in crayfish and mitigate WSSV-induced pathological damage in key tissues such as the hepatopancreas, intestines, and gills. These results provide compelling evidence of the compound’s antiviral efficacy. However, the precise antiviral mechanisms and the primary bioactive constituents remain to be fully explored. Modern pharmacological research has revealed that baicalin, a key flavonoid in SB, can modulate oxidative stress-induced inflammation by regulating the PI3K/Akt/NRF2 axis and nuclear factor-кB (NF-кB) signaling, and exerts antiviral effects against IVA, RSV, HV, and CVB3 via modulation of JAK/STAT, TLRs, and NF-кB cascades.22,23 Besides, Rutin, a major compound in FSI, has demonstrated antioxidant and anti-inflammatory effects in zebrafish models and neuroprotective effects against ischemic stroke under strict dosage control.24 Meanwhile, PV-derived polysaccharide PVE30 was found to effectively inhibit herpes simplex virus (HSV) by preventing TLR-mediated NF-кB activation.25 The ursolic acid component of PV was found to demonstrate strong activity against IHNV in rainbow trout, enhancing relative survival by 18.9%.26 Collectively, the established anti-inflammatory and antiviral properties of these herbs support the reliability and mechanistic plausibility of our research findings. Additionally, RT-qPCR results showed that the compound formulation, composed of SB, FSI, and PV, increased the expression levels of several innate immune-related genes in crayfish. Among these, the calcium-dependent (C-type) lectin (PcLT), a typical pattern recognition receptor (PRR), plays a pivotal role in immune surveillance by recognizing specific carbohydrate structures via its carbohydrate recognition domain (CRD). PcLT is known to activate the prophenoloxidase (proPO) system and mediate cellular immunity by promoting hemocyte adhesion, ultimately contributing to the clearance of invading viruses.27–29 Antilipopolysaccharide factors ALF1 and ALF2, members of the antimicrobial peptides (AMPs) family, recognize bacterial lipopolysaccharides and exert antimicrobial and antiviral effects membrane disruption of pathogens. In crustaceans, WSSV infection has been shown to trigger the Toll-like receptor-mediated NF-кB signaling pathway, which in turn regulates antimicrobial peptides (AMPs) gene expression.30,31 Furthermore, WSSV infection triggers ROS overproduction, leading to oxidative stress in shrimp. Antioxidant enzymes, exemplified by glutathione S-transferase (GST), superoxide dismutase (SOD), and catalase (CAT), are indispensable for sustaining the intracellular redox balance. These enzymes protect cells from oxidative damage during viral invasion, thereby enhancing the disease resistance of crayfish.32,33 Building upon the above findings, it can be inferred that SB, FSI, and PV can enhance the expression of these innate immune genes to exert antiviral effects, thereby boosting the immune response of crayfish.
The intestinal microbiota plays a pivotal role in host metabolic processes, nutrient uptake, and immune modulation. Its composition and diversity are dynamic and affected by various internal and external factors throughout the host’s life cycle.34–38 Accumulating evidence suggests that viral invasion can disrupt the equilibrium of the intestinal microbiota, impairing its physiological functions.39,40 In aquatic animals, several fish viruses have been shown to reduce gut microbial diversity and abundance. For instance, grass carp reovirus and koi herpesvirus type II infections were found to significantly alter the gut microbial composition, structure, and diversity in grass carp and crucian carp, respectively.41,42 In this research, analysis of gut microbial composition and diversity in crayfish revealed that both WSSV infection and drug treatment influenced microbial richness and evenness. At the phylum level, Proteobacteria was the prevalent phylum in all experimental groups, consistent with previous findings in other aquaculture animals.43 Notably, WSSV infection resulted in a substantial increase in Proteobacteria abundance in the CHG group. Elevated levels of Proteobacteria have also been reported in diseased crayfish, and excessive proliferation of this phylum has been associated with impaired digestive function and inflammation.44–46 In the present study, treatment with the compound drug significantly reduced Proteobacteria abundance in the TRE and DTG groups, suggesting that this formulation could alleviate gut dysbiosis and reduce microbial-induced intestinal injury. This is further substantiated by previous studies showing that rutin, a key flavonoid in FSI, can partially restore gut microbiota imbalance in murine models of colitis.47 In addition to changes in Proteobacteria, notable shifts were observed in the relative abundance of Firmicutes and Bacteroidetes, two major phyla that are commonly used as indicators of intestinal homeostasis in mammals. In contrast to the CON group, the CHG group displayed a marked decline in Firmicutes and an increase in Bacteroidetes. Interestingly, the DTG group exhibited an increase in both phyla, indicating a potential recovery toward microbial balance. In mammals, the F/B ratio is often considered a health indicator for the gut, with a reduced F/B ratio being linked to intestinal inflammation and dysbiosis.48 In this study, WSSV infection led to a decreased F/B ratio across all infected groups, with the most pronounced reduction observed in the CHG group. This finding aligned with the intestinal epithelial damage observed in histopathological sections, further suggesting that WSSV infection induces gut inflammation and that the compound drug may play a protective role in conserving intestinal integrity.
At the genus level, the Bacteroides genus was identified as a core shared among all four groups, suggesting its role as a resident microbiota in crayfish. This genus has been linked to human gut inflammation.49 In this study, the abundance of Bacteroides was consistently above 5% in all groups, with the CHG group displaying a markedly elevated abundance compared to the other groups. A similar increase in Bacteroides abundance has been documented in grouper following iridovirus infection.50 Based on the above findings, it can be inferred that WSSV infection contributed to the observed rise in Bacteroides abundance. Pseudomonas, although not typically a core member of the gut microbiota, may disrupt microbial balance and diversity in diseased groups, potentially leading to abnormal immune responses and disease.51 Accordingly, the comparatively high abundance of Pseudomonas in the CHG group is due to WSSV infection. In sea urchins, Pseudomonas abundance was found to be significantly increased in individuals suffering from red spot disease, which supports our findings. Most studies suggest that flavonoids help maintain gut microbiota stability and foster the proliferation of health-promoting microbes.52,53 Shewanella, a genus belonging to the Proteobacteria phylum, has been implicated in disease outbreaks in crayfish under specific conditions. However, previous studies have demonstrated that adding mulberry leaf flavonoids to the feed of Litopenaeus vannamei promotes the proliferation of Shewanella.54 In the present study, relative to the CHG group, the abundance of this genus in the TRE group was substantially diminished, indicating that the flavonoid-rich SB and FSI effectively suppressed the abundance of Shewanella to preserve the intestinal health of crayfish.
The metabolome is a direct reflection of the functional crosstalk between the intestinal microbes and their host, with the two systems functioning in a complementary manner.55 Correlation analyses integrating 16S sequencing and metabolomics have shown that the genus Bacteroides is positively correlated with numerous metabolites involved in amino acid, fatty acid, and lipid metabolism.56 In Atlantic salmon, replacing fructooligosaccharides with galactooligosaccharides significantly elevated metabolites associated with phospholipid, fatty acid, carnitine, and sphingolipid metabolism, and these changes were strongly associated with shifts in the abundance of Firmicutes and Actinobacteria.57 During this study, non-targeted metabolomics analysis found that WSSV infection and compound drug treatment not only affected the abundance of the gut microbiota in crayfish but also caused marked changes in the levels of gut-derived metabolites, highlighting the close interplay between microbial structure and host metabolic profiles.
To demonstrate the effects of the compound on gut metabolites in crayfish infected with WSSV, as well as the relationship between specific metabolite alterations and associated changes in gut microbiota, KEGG pathway enrichment analysis was conducted on the differentially expressed metabolites found in the CHG and TRE groups. Overall, 20 significantly enriched metabolic pathways were identified, including cholesterol metabolism, glycerophospholipid metabolism, arachidonic acid metabolism, and steroid metabolism. Relative to the CHG group, the metabolism of glycerophospholipids was significantly upregulated in the TRE group. Importantly, our correlation analysis revealed that this metabolic upregulation showed significant positive correlations with herb-promoted beneficial bacteria (such as Anaerorhabdus and Propionivibrio) and negative correlations with WSSV-induced harmful bacteria (such as Shewanella and Bacteroides-H) (Appendix Figure 2). Glycerophospholipids are major components of cell membranes, maintaining cell integrity and fluidity, as well as participating in intracellular signaling and material transport.58,59 Their upregulated metabolism helps enhance cell membrane stability and prevents WSSV invasion. Interestingly, an upregulation in arachidonic acid metabolism, a metabolite of glycerophospholipids, was also observed in the TRE group. Correlation analysis further confirmed that the accumulation of arachidonic acid was closely associated with the proliferation of beneficial bacteria. Arachidonic acid and its derivatives are known to be key mediators in immune regulation and lipid metabolic homeostasis,60,61 and can activate the NF-κB signaling pathway in crustaceans, enhancing antimicrobial peptide expression. Therefore, the herbal compound may drive the activation of the glycerophospholipid-arachidonic acid metabolic axis by promoting the proliferation of specific beneficial bacteria, thereby enhancing the host’s antiviral immune response.
In addition to the aforementioned pathways, the TRE group also exhibited significant upregulation in several key metabolic pathways, including steroid and steroid hormone biosynthesis, pantothenate and CoA biosynthesis, and pyrimidine metabolism. Notably, key metabolites in these pathways also showed positive correlations with beneficial bacteria, forming a coordinated microbiota remodeling that drives metabolic reprogramming and leads to an enhanced immune defense network. Steroid hormones, particularly glucocorticoids, are renowned for their potent anti-inflammatory properties, which they exert by suppressing the activity of inflammatory cells and decreasing the production of pro-inflammatory cytokines.62 Pantothenate is a key precursor for the synthesis of Coenzyme A (CoA), which plays a central role not only in energy provision but also in regulating glutathione metabolism to enhance antioxidant defense mechanisms.63 Pyrimidine metabolites are core components of nucleotides and their derivatives, participating in glycogen synthesis, phospholipid metabolism, and cell signaling cascades.64–66 Furthermore, certain pyrimidine metabolites such as cytosine phosphoguanine (CpG) can enhance antiviral immune responses by modulating T-cell activity.67
Overall, this study revealed that the compound drug composed of SB, FSI, and PV in a ratio of 2:2:1 could effectively reduce the viral load in crayfish following WSSV infection, promote the expression of innate immunity-associated genes, and alleviate tissue damage in hepatopancreas, intestines, and gills. An integrated analysis combining 16S rRNA sequencing and non-targeted metabolomics further revealed that this compound modulated the gut microbiota composition and altered key metabolites, thereby mitigating WSSV-induced gut dysbiosis and metabolic disturbances. Collectively, these findings indicate that the herbal compound exerts a significant anti-WSSV effect on crayfish, providing a solid foundation for its potential application in the management of viral diseases in aquaculture.
ACKNOWLEDGMENTS
This work was funded by Central Public-interest Scientific Institution Basal Research Fund (No. YFI20240403) and Central Public-interest Scientific Institution Basal Research Fund, CAFS (No. 2023TD46).
AUTHORS’ CONTRIBUTION
Conceptualization: Yun Feng (Equal), Zhenyu Huang (Equal). Data curation: Yun Feng (Equal), Zhenyu Huang (Equal), Yuding Fan (Equal). Formal Analysis: Yun Feng (Equal), Yong Zhou (Equal). Investigation: Yun Feng (Equal), Zhenyu Huang (Equal), Mingyang Xue (Equal). Methodology: Yun Feng (Equal), Zhenyu Huang (Equal), Mingyang Xue (Equal). Validation: Yun Feng (Equal), Zhenyu Huang (Equal), Yong Zhou (Equal). Visualization: Yun Feng (Equal), Zhenyu Huang (Equal), Yuding Fan (Equal). Writing – original draft: Yun Feng (Equal). Resources: Zhenyu Huang (Equal), Yuding Fan (Equal), Yong Zhou (Equal). Writing – review & editing: Zhenyu Huang (Equal). Software: Chen Xu (Equal), Yan Meng (Equal), Nan Jiang (Equal), Juan Tian (Equal). Supervision: Yan Meng (Equal), Nan Jiang (Equal), Yiqun Li (Equal), Wenzhi Liu (Equal), Xin Ren (Equal). Funding acquisition: Lin Zhang (Equal), Yong Zhou (Equal). Project administration: Lin Zhang (Equal), Ya Zheng (Equal), Yong Zhou (Equal).
COMPETING OF INTEREST
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
ETHICAL CONDUCT APPROVAL
All animal experiments were conducted in compliance with the Ethical Inspection for Animal Experimentation.
INFORMED CONSENT STATEMENT
All authors and institutions have confirmed this manuscript for publication.

_experimental_design_schematic_showing_tre.tiff)
_-diversity_indices__c.tiff)
_heatmap_of_microbial_composition_a.tiff)
_principal_component_analysis.tiff)
_metabolite_identif.tiff)
_staining._(.tiff)
_experimental_design_schematic_showing_tre.tiff)
_-diversity_indices__c.tiff)
_heatmap_of_microbial_composition_a.tiff)
_principal_component_analysis.tiff)
_metabolite_identif.tiff)
_staining._(.tiff)