1. Introduction
Redclaw crayfish (Cherax quadricarinatus) is a freshwater species that inhabits streams, rivers, and lakes in Australia. It has the advantages of environmental tolerance, large body size, strong stress resistance, delicious meat quality, and rich nutrition. Its tail meat yield is five times that of ordinary crayfish. Therefore, the redclaw crayfish has significant economic value and has been regarded as one of the most suitable and popular aquaculture species in the world.1,2 The redclaw crayfish grows rapidly and can reach commercial specifications (50-100 grams) within 7 months in a suitable environment.3,4 However, with the rapid development of aquaculture in recent years, the deterioration of the water environment and the increase in aquaculture density have led to frequent aquatic animal diseases, which have seriously affected the output and economic benefits of aquaculture.5,6 In order to control the spread of pathogens, disinfectants are widely used in aquaculture.7
Formaldehyde (HCHO) is one of the most widely used fungicides in aquaculture.8,9 In aquaculture, formaldehyde is mainly used to prevent and treat bacterial, parasitic, and fungal infections, especially in high-density aquaculture environments, helping to reduce the incidence of diseases.10,11 However, the widespread use of formaldehyde in the aquaculture process can lead to environmental degradation and may have negative effects on treated fish.12
In recent years, with the development of microbiology and immunology, people have gradually realized that gut microbiota not only plays a key role in maintaining host health, but also plays an important role in defending against pathogen infection.13 The role of gut microbiota in pathogen infection is multi-fold, involving colonization resistance, immune regulation, production of metabolites, and direct inhibition of pathogens.14 The gut microbiota restricts the growth and colonization of pathogens by competing for nutrients and colonization sites.15 In addition, the growth of pathogens can be directly inhibited by gut microbiota through the production of metabolites such as antimicrobial peptides and short-chain fatty acids.16–18 Studies have shown that a stable gut microbiota can promote the growth and development of aquatic animals while enhancing their resistance to pathogens.19 On the contrary, an imbalance in the gut microbiota may increase the risk of disease.20,21 Although formaldehyde, as a widely used disinfectant in aquaculture, has been proven to have a remarkable bactericidal effect, there is still a lack of systematic research on its influence mechanism and potential ecological risk on the intestinal microecosystem of redclaw crayfish.
At present, the regulatory effect of formaldehyde exposure on the composition and function of crustacean intestinal flora and the interaction between host and microorganism is unclear, especially the mechanism of its influence on the intestinal microecological balance of red crayfish, an important economic species, needs to be clarified. However, there are few studies on the effects of formaldehyde on the gut microbiota of redclaw crayfish. Therefore, this study intends to analyze the changes in gut microbiota of redclaw crayfish exposed to formaldehyde through metagenomic sequencing technology and explore its potential impact on gut microbial homeostasis and host health. To deeply study the microbial adaptation mechanism of aquatic organisms to exogenous chemical substances, reveal their ecological impact and regulation mechanism, and thus provide scientific theoretical support for the development of environmentally friendly aquaculture technology.
2. Materials and methods
2.1. Animals and experimental design
The experiments were performed at the Zhejiang Institute of Freshwater Fisheries, Huzhou, China. All experimental specimens of redclaw crayfish were obtained from a commercial crayfish farm in Wencheng County, Wenzhou City, Zhejiang Province, China. A total of twenty redclaw crayfish (body weight: 50 ± 0.15 g) at 10 months of age were used for the experiment. The experiment was conducted in a 100-liter polyethylene plastic barrel. The culture tank water was maintained at a temperature of 26 ± 0.5 ◦C, salinity at 3‰, and pH at 7.2 ± 0.2. Before the experiment, redclaw crayfish were acclimatized in the containers for 7 days and were fed commercial pellet feed (Guangdong Hengxing, China) twice daily at 8:00 am and 5:00 pm until satiation was achieved. Following a 7-day acclimatization period, the formaldehyde exposure experiment was initiated.
A one-hour prophylactic static bath with formaldehyde was performed, in which healthy redclaw crayfish were exposed to formaldehyde solutions at concentrations of 0 ppm (control group, CK, n = 10) and 100 ppm (treatment group, IN, n = 10). At one hour of exposure, intestinal contents were collected from twenty redclaw crayfish in both the control (n = 10) and treatment groups, and 100 ppm (n = 10).
One part of the sample was taken for microbiome analysis and stored at − 80 ◦C. The samples from the control group were processed in the same manner as those from the treatment group, with five replicates performed for the microbiome analyses. The study procedures were approved by the Ethics Committee of the Animals Research Centre of Zhejiang Province.
2.2. Genomic DNA extraction and sequencing
We extracted genomic DNA from 200 mg intestinal contents using an optimized CTAB protocol. Samples were homogenized in CTAB buffer using a Precellys 24 homogenizer, followed by proteinase K digestion. DNA quality was verified by electrophoresis and spectrophotometry. Qualified DNA libraries were sequenced by Illumina Novaseq high-throughput sequencing platform, and the raw data obtained from sequencing was used for bioinformatic analysis.
2.3. Bioinformatics and statistical analysis
In order to ensure the reliability of data, the Raw sequencing data needs to be preprocessed using Kneaddata software. FastQC is used to detect the rationality and effect of quality control (Langmead and Salzberg22; Martin23; Robert et al., 2011). We performed taxonomic profiling of microbial communities using Kraken2 (version 2.1.2), a k-mer-based classification tool, with a custom-built microbial database constructed by screening bacterial, fungal, archaeal, and viral sequences from NCBI’s NT nucleic acid database and RefSeq whole genome database. Kraken2 analysis was conducted with a confidence threshold of 0.2 (–confidence 0.2) to optimize sensitivity for species detection. The resulting classifications were then processed with Bracken to estimate species-level relative abundances using default parameters, which probabilistically re-distributes reads across the taxonomic tree to provide more accurate abundance measurements. This combined approach leverages Kraken2’s rapid k-mer matching for initial classification and Bracken’s Bayesian re-estimation method to generate refined quantitative profiles of microbial community composition. The clean reads after quality control and de-host were used to blast to database (Uniref90) using Humann2 software (based on Diamond), and the annotation information and relative abundance table from each functional database were obtained according to the corresponding relationship between Uniref90 ID and each database.24–27 Lefse Biomarker and Dunntest analysis performed to excavate differences in species composition and functional composition between samples.28
3. Results
3.1. OTUs and diversity analysis of gut flora
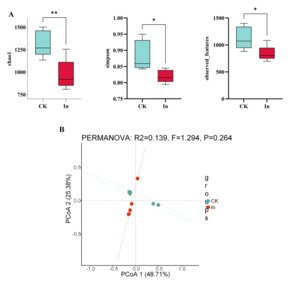
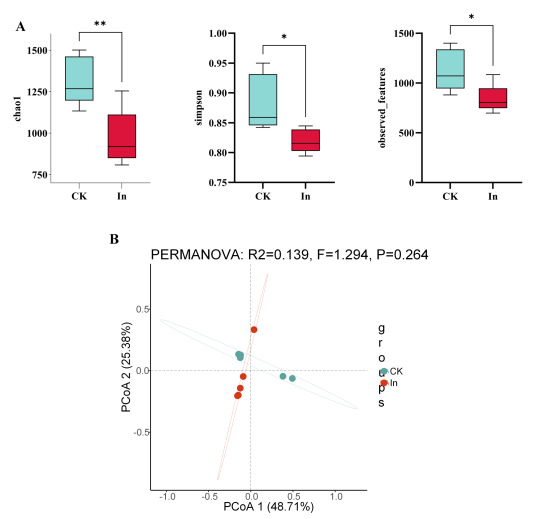
Metagenomic sequencing gave a mean 27,080,833 clean reads per sample for CK and 24,754,015 clean reads per sample for In. Based on 97% similarity, 2433 and 1802 OTUs were separately acquired from CK and In groups, respectively. We statistically analyzed the effects of formaldehyde on intestinal microbiota diversity in redclaw crayfish using Independent Samples t-tests for diversity indices, with subsequent False Discovery Rate (FDR) adjustment to account for multiple comparisons. Compared with the CK group, the Chao1 and Simpson index of the In group were significantly lower than those of the CK group (p<0.05) (Fig. 1A). PCoA analysis showed that the gut microbiota of the In group deviated from the control group (Fig. 1B). In summary, formaldehyde exposure has a significant effect on the diversity of redclaw crayfish gut microbiota.
3.2. Results of annotation of gut bacterial levels in redclaw crayfish
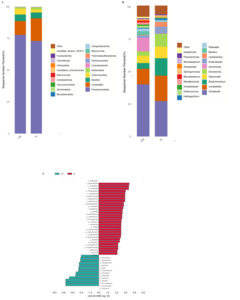
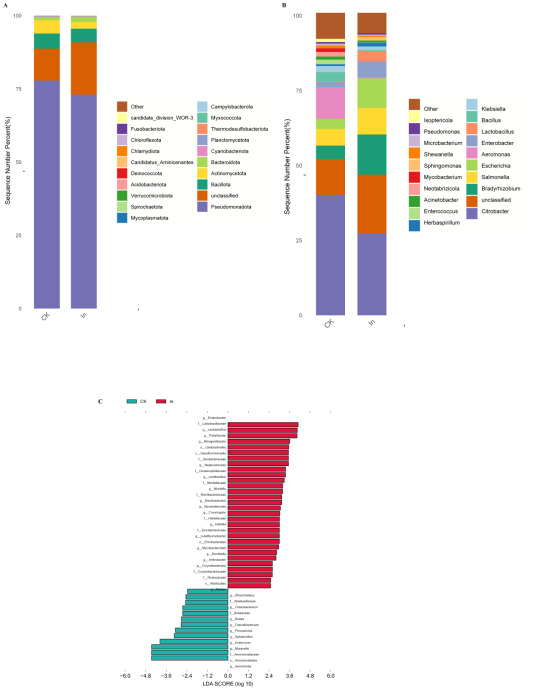
In this study, Pseudomonadota, Bacillota, Actinomycetota, Bacteroidota and Cyanobacteriota were the dominant phyla in the gut of redclaw crayfish (Fig. 2A). The dominant genera in redclaw crayfish were Citrobacter, Bradyrhizobium, Salmonella, Escherichia and Aeromonas (Fig. 2B).
An LEfSe determination (LDA score > 2) was performed to identify the key differential taxa in the redclaw crayfish gut bacterial (Fig. 2C, Table S1). Formaldehyde exposure caused decreases in the relative abundance of some bacterial families including Lactobacillaceae, Brevibacteriaceae, Oceanospirillaceae and Moritellaceae (Table S1), and in some bacterial genera including Enterobacter, Lactobacillus, Polaribacter, Nitrogeniibacter and Neptunomonas (Table S2). The relative abundance of Aeromonas, Moraxella, Acidovorax, Sphaerotilus and Phocaeicola increased after formaldehyde exposure.
3.3. Results of annotation of gut fungal levels in redclaw crayfish
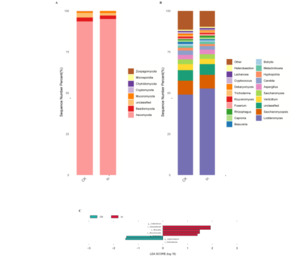
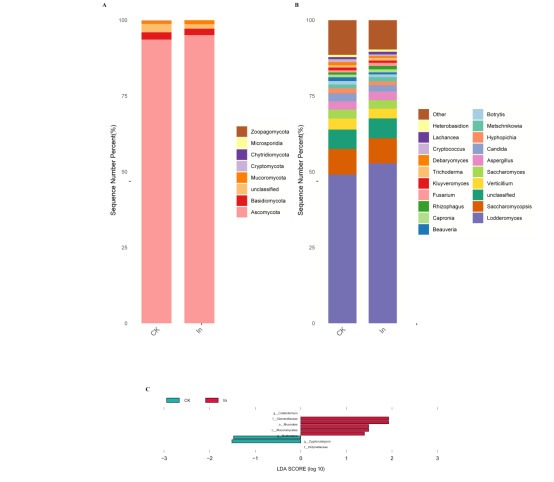
The results of this study showed that the dominant phyla in the gut of redclaw crayfish were Ascomycota, Basidiomycota, Chytridiomycota, and Microsporidia (Fig. 3A), and the dominant genera were Lodderomyces, Saccharomycopsis, Verticillium, Saccharomyces and Aspergillus (Fig. 3B).
LEfSe analysis showed that the relative abundance of Glomerulaceae and Arthrodermataceae decreased, while the relative abundance of Didymellaceae increased in the In group at the family level compared with the control group (Fig. 3C Table S3). At the genus level, the relative abundance of Colletotrichum and Neohortaea decreased, while that of Zygotorulaspora increased in the In group compared with the control group (Fig. 3C, Table S4).
3.4. Results of annotation of gut archaea levels in redclaw crayfish
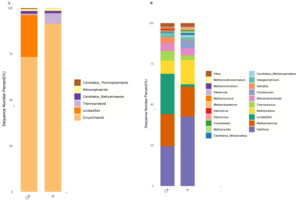
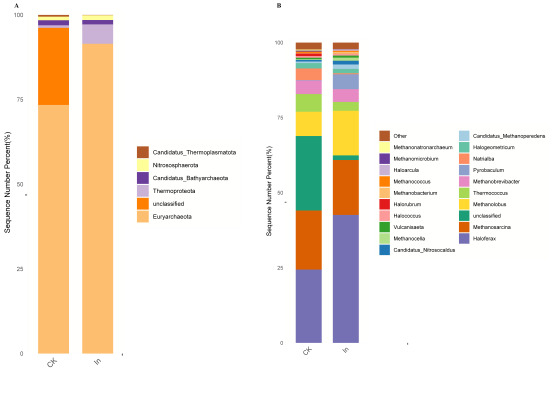
The results of this study showed that candidatus, bathyarchaeota, candidatus thermoplasmatota, euryarchaeota, nitrososphaerota and thermoproteota were the dominant bacteria in the redclaw crayfish gut (Fig. 4A). Haloferax, methanosarcina, methanolobus, thermococcus and methanobrevibacter were the dominant genera in the fish gut. It is worth noting that haloarcula was unique to the In group (Fig. 4B).
3.5. Analysis of differential metabolic pathways between groups
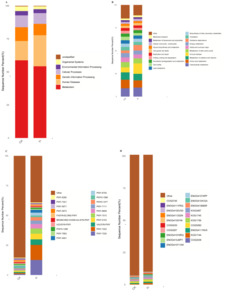
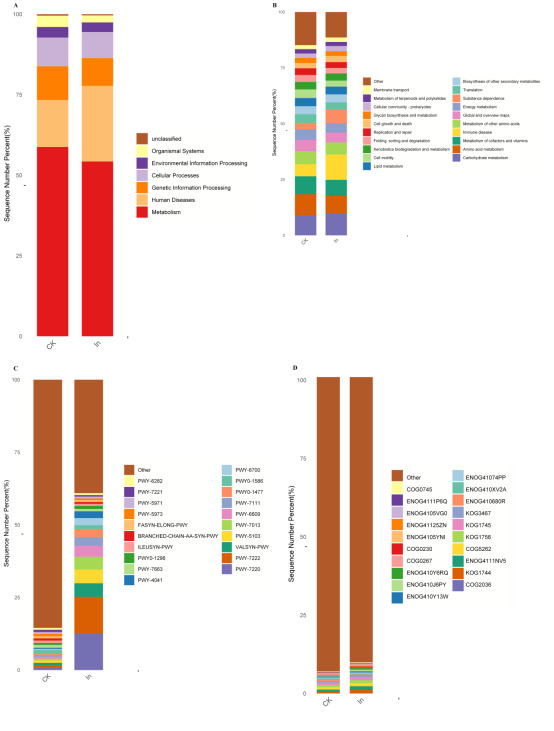
KEGG database statistics (Fig. 5A), the number of genes annotated by metabolism in the CK and In groups was the highest in the first level classification. At the secondary level (Fig. 5B), the CK group had the highest number of genes annotated in the amino acid metabolism of the gut microbiota, followed by carbohydrate metabolism, metabolism of cofactors and vitamins, metabolism of other amino acids, and immune disease. In the In group, the number of genes annotated in immune disease was the highest, followed by carbohydrate metabolism, amino acid metabolism, metabolism of cofactors and vitamins, and substance dependence.
According to the annotation results of MetaCyc database, relative abundance statistics of each channel in each experimental group were drawn (Fig. 5C). In the CK group, the number of genes annotated in PWY0-1586 was the largest, followed by PWY-7663、VALSYN-PWY and PWY-5103. In the In group, the number of genes annotated in PWY-7220 was the largest, followed by PWY-7222、PWY-5103 and VALSYN-PWY.
Eggnog database statistics (Fig. 5D), the number of annotated genes in the intestinal microbiota COG2036 in the CK and In groups were the highest, followed by kog1744, enog4111nv5, cog5262 and kog1756.
4. Discussion
As a highly effective broad-spectrum disinfectant, formaldehyde is widely used in aquaculture to control infections by pathogens such as fungi, bacteria and parasites on the surface of fish. However, studies have shown that formaldehyde exposure can cause a variety of pathological and physiological symptoms in farmed fish, but its underlying toxicity mechanism has not yet been fully elucidated.29–31 Given the key regulatory role of the gut microbiota in maintaining host health, we speculate that the interaction between formaldehyde and the gut microbiome may be one of the important mechanisms of its toxic effects.32
Intestinal microbiota plays a crucial role in maintaining host immune function, regulating metabolic homeostasis, and providing resistance against pathogenic infections. Recently, it has been reported that the changes in intestinal microflora are related to the pathogenesis of many inflammatory diseases and infections.33,34 In studies on zebrafish, goats, mouse and Plagiodera versicolora, it was found that gut microflora can be used as a physical barrier to inhibit pathogens from invading the host gut and improve the host immunity.35–37
In recent years, research on the gut microbiota of redclaw crayfish has made significant progress, but research on the effects of formaldehyde exposure on the gut microbiota of this species is still relatively scarce. Therefore, this study used metagenomics technology to systematically elucidate the regulatory mechanism of formaldehyde exposure on the structure and functional characteristics of the intestinal microbial community of red claw crayfish, focusing on the dynamic changes in the community composition of the intestinal flora under formaldehyde stress, the functional response of key metabolic pathways, and the potential environmental adaptation mechanism. The results of this study showed that formaldehyde exposure significantly changed the composition and diversity characteristics of the intestinal microbial community of red claw crayfish. These findings provide new insights into the ecotoxicological effects of formaldehyde exposure on crustaceans.
This study used metagenomic sequencing technology to systematically analyze the composition characteristics of the intestinal microbiome of redclaw crayfish from three dimensions: bacterial community structure, archaeal composition characteristics, and fungal diversity, and evaluated the mechanism of formaldehyde exposure on the structure, function, and diversity of its intestinal flora. The characterization of redclaw crayfish intestinal microbiota revealed Pseudomonadota, Bacillota, Actinomycetota, Bacteroidota and Cyanobacteriota as the predominant phylum.38 The characterization of redclaw crayfish Intestinal microbiota communities revealed Citrobacter, Bradyrhizobium, Salmonella, Escherichia and Aeromonas as the predominant genera. LEfSe analysis results showed that after formaldehyde exposure, the relative abundance of redclaw crayfish Lactobacillus Nitrogeniibacter, Aeromonas and Phocaeicola in the intestine had changed significantly. Lactobacillus had broad-spectrum antibacterial activity, which can inhibit many pathogenic bacteria and improve the survival rate of shrimp.39 Nitrogenibacter is a nitrifying bacterium, which was used in a bioremediation system to control the concentration of nitrogen compounds in shrimp ponds. For example, the use of nitrifying bacteria in aquaculture can significantly reduce the concentration of ammonia and nitrite in shrimp ponds and improve the survival rate of shrimp.40 Aeromonas was a gram-negative bacterium, which had a significant negative impact on aquaculture and can infect many animals, including sea bass, Crucian Carp, alligator, Yangtze finless porpoise, Trionyx sinensis and sheep etc.41–47 Phocaeicola is a gram-negative, strictly anaerobic bacterium that widely exists in animal intestines and has remarkable functions in digesting complex carbohydrates and producing short-chain fatty acids (SCFAs).48 For example, Phocaeicola vulgatus was one of the most abundant bacteria in human feces, which can decompose dietary fiber and produce SCFAs such as acetic acid, propionic acid, and butyric acid, which were beneficial to intestinal health and general health.49 Also, Phocaeicola interacts with other microorganisms in the intestine to inhibit the growth of pathogenic bacteria through competition, thus maintaining intestinal homeostasis. The changes of Phocaeicola abundance in intestinal microbiota were related to many diseases, such as inflammatory bowel disease (IBD) and irritable bowel syndrome (IBS).50,51 Therefore, Phocaeicola played an important ecological and physiological role in intestinal microflora, and its research was helpful in developing strategies to promote intestinal health and treat related diseases. The significant change of relative abundance of Lactobacillus, Nitrogeniibacter, Aeromonas and Phocaeicola may be related to the damage of intestinal cells. However, its ecological function in crustaceans has not yet been clarified, and the specific function of this bacterium still needs to be further clarified through crustacean-specific studies (such as metagenomic analysis and in vitro verification). This study showed that acute formaldehyde exposure changed the composition of the intestinal flora of redclaw crayfish and may affect host health through microecological imbalance. However, due to the lack of simultaneous detection of host physiological indicators, the direct correlation between this change in microbiome structure and host physiological function still needs further verification. Future studies need to further verify its biological impact mechanism by integrating multi-omics analysis (such as transcriptome, proteome) and host function experiments (such as immune response assays and pathogen challenge tests).
Afterwards, from the perspective of fungi, the characterization of redclaw crayfish gut flora communities revealed Ascomycota, Basidiomycota, Chytridiomycota and Microsporidia as the predominant phylum.38 This was consistent with the analysis results of intestinal fungi by Xu.52 Wang et al. detected Colletotrichum in the intestines of horses, and found that the fungal flora in the intestines changed obviously after natural infection with Cryptosporidium.53 The results were consistent with our research. Zygotorulaspora is a yeast belonging to Ascomycota, which was significantly increased in the gut microflora of mice with micronutrient deficiency in early life.54 Therefore, changes in the intestinal flora structure caused by formaldehyde exposure may disrupt the intestinal microecological balance, lead to impaired intestinal barrier function of the host, and affect the normal function of its immune regulation mechanism.
Archaea played an important role in intestinal microflora, and the diversity of gut archaea was related to the development of inflammation.55 Our analysis of archaea revealed that candidatus_bathyarchaeota, candidatus_thermoplasmatota and euryarchaeota were the dominant phylum in redclaw crayfish intestine, Haloferax, Methanosarcina, Methanolobus and Thermococcus was the dominant genus in redclaw crayfish intestine. It was worth noting that Haloarcula was unique to group In. Borjian56 and others found that Haloarcula has a unique metabolic pathway and enzyme system. For example, the methanesulfonic acid cycle in Haloarcula hispanica was one of its unique metabolic pathways, which involves many key enzymes, such as succinyl-CoA:mesaconate CoA-transferase and mesaconyl-CoA hydratase.56 However, although archaea have been shown to have unique metabolic characteristics (such as the methane sulfonate cycle pathway and related enzyme systems),56 as a non-dominant component of the intestinal microbiome, their functional characterization in the intestinal ecosystem of redclaw crayfish remains unclear.
In this study, an experimental design of 100 ppm formaldehyde exposure for 1 h was used to evaluate the effects of acute exposure conditions on the intestinal flora of redclaw crayfish. Although this scheme provides an important reference for understanding the potential toxic effects of formaldehyde, we also recognize that there may be more complex application scenarios in actual aquaculture production. In conventional farming, formaldehyde is often used in a flexible treatment scheme, and environmental conditions may also have a regulatory effect on the treatment effect. In future studies, more attention should be paid to the comprehensive evaluation of formaldehyde gradient concentrations and chronic exposure on the ecotoxic effects of redclaw crayfish.
5. Conclusion
This study systematically revealed the significant effects of formaldehyde exposure on the structure and functional characteristics of the intestinal microbial community of redclaw crayfish through metagenomic sequencing technology. The results showed that formaldehyde exposure significantly reduced the alpha diversity index of the intestinal microorganisms of redclaw crayfish (Chao1, Simpson and observed features index, p<0.05) and changed its community composition. In terms of bacterial communities, formaldehyde exposure led to a significant decrease in the relative abundance of beneficial bacteria such as Lactobacillus and Nitrogeniibacter, while a significant increase in the relative abundance of conditional pathogens such as Aeromonas. Fungal community analysis showed that Ascomycota and Basidiomycota were the dominant phyla, among which the relative abundance of Zygotorulaspora increased significantly. In the archaeal community, Candidatus_Bathyarchaeota and Candidatus_Thermoplasmatota were the dominant phyla, and Haloarcula was a genus unique to the exposure group. Metabolic pathway analysis showed that formaldehyde exposure affected key metabolic pathways such as amino acid metabolism and carbohydrate metabolism, and increased the number of annotations of immune-related genes. These results suggest that formaldehyde exposure may affect the host’s intestinal microecological balance by altering the structure and function of the intestinal flora. However, the specific mechanism of immune function regulation still requires further research and verification. This study provides new insights into the ecotoxicological effects of formaldehyde on crustaceans and has important guiding significance for the formulation of safe use standards for formaldehyde in aquaculture.
Acknowledgments
The authors declare that they have not recognized competing financial interests or personal connections that could influence the work presented in this paper. This study was supported by the Seed Capital for Innovation and Entrepreneurship of Wencheng County (GrantNO.2023NKY05).
Authors’ Contribution – CRediT
Writing - original draft preparation: Haiheng Wang, Yanyun Liu and Wenwu Zou; Writing - review and editing: Shun Cheng; Funding acquisition: Haiheng Wang; Formal analysis and investigation: Zhilong Chen and Qinyi Xiong.
Competing of Interest – COPE
The authors declare that they have no recognized competing financial interests or personal connections that could influence the work presented in this paper.
Ethical Conduct Approval – IACUC
The study procedures were approved by the Ethics Committee of Animals Research Centre of Zhejiang Province.
Informed Consent Statement
All authors and institutions have confirmed this manuscript for publication.
Data Availability Statement
All are available upon reasonable request.