Introduction
Acute hepatopancreatic necrosis disease (AHPND), also known as Early mortality syndrome, is one of the most severe problems in aquaculture. It is a highly lethal bacterial infection that affects farming shrimps, primarily effecting the white leg shrimp Litopenaeus vannamei, and the giant tiger prawn shrimp Penaeus monodon, resulting infected shrimps present a pale and atrophied hepatopancreas together with an empty stomach and mid-gut.1–3 What is particularly worrisome is that the mortality of up to 100% in shrimps within 30-35 days of stocking.3,4 The shrimps effected by AHPND exhibit distinctive features, including lethargy, anorexia, slow growth, empty digestive tract and a pale to white hepatopancreas.5,6
Historically, since AHPND was first documented in China in 2009, it has marked the beginning of widespread outbreaks of disease across global farming shrimps led to a significant threat to aquaculture.7,8 Following the first document, AHPND was reported in Vietnam in 2010, Thailand in 2011 and 2012.2,9 The emergence of AHPND in Vietnam, and Thailand marked an outbreak in other parts in Southeast Asian region. The disease of AHPND spread throughout the Americas: Mexico in 2013, South America in 2016, and America in 2019.6,10 Over the years, the disease’s quick global spread highlighted how disruptive it could be to shrimp aquaculture globally.
The current study aimed to thoroughly provide a comprehensive analysis of the genetic variability of Vibrio parahaemolyticus strains, specially focusing on the PirABVP toxins. Additionally, this research also seeks to provide molecular insights by systematically comparing genetic variations across pathogenic, non-pathogenic, and mutant strains. this research also seeks to point out the PirABVP pathogenicity through the exploration of the molecular mechanisms underlying these toxins, the similarity of their structure to Cry proteins, and their contribution to virulence through horizontal gene transfer. These summaries will contribute to the development of targeted diagnostic tools, therapeutic interventions, and preventive strategies to mitigate the impact of AHPND on shrimp aquaculture.
Plasmid pVA1 - associated genetic factors driving AHPND pathogenesis
AHPND-infected shrimps exhibit the symptoms of lethargic and display erratic swimming behavior, finally leading to the dysregulation and damage of hepatopancreas, following by rapid mortality.3,6 However, these reported signs are nonspecific and common for other shrimp diseases. Thus, making it challenging to diagnose AHPND solely on the basis of physical symptoms. Given that these signs overlap with other diseases, it is crucial to identify bacterial causative to confirm AHPND. Earlier report in 2013 revealed that AHPND is caused by virulent strains of Vibrio parahaemolyticus, which widely infect Penaeus monodon, Litopenaeus vannamei, namely as AHPND-causing Vibrio parahaemolyticus (VPAHPND).2,5,6,11
V. parahaemolyticus, a gram-negative, rod shape and halophilic bacteria, was first identified in 1950 by Tsunesaburo Fujino. It was confirmed as a causative agent of food born disease.1,3,12 Most of harmless V. parahaemolyticus are isolated from aqueous environment due to the lack of two pathogenic genes: tdh (thermostable direct hemolysis), and trh (TDH-related hemolysin), which function as hemolysis and cytotoxicity activity in the host cell, and other virulent factors, systems, such as QS (quorum sensing), adhesion, type 3 secretion system (T3SS1, T3SS2), type 6 secretion system (T6SS1, T6SS2) systems or extrachromosomal pathogenic plasmids.3,13–16 Whereas some of the V. parahaemolyticus strains gain virulent genes, and systems have been reported to be pathogenic to animal and human.13,14,16 Apart from above, VPAHPND has been currently reported as aquatic zoonotic pathogen that can cause vibriosis in many fish and shellfish species, especially shrimps, because VPAHPND harbor two Photorhabdus insect-related (Pir) toxin genes A and B (PirAVP (ORF51), and PirB (ORF51)) locating at a 70 kps-length exogenous chomosomal plasmid – pVA1, contains about ~90 open-reading frames (ORFs), which are not found in non-pathogenic strains.1,6,16,17 The removal of pVA1 or selective knockdown of Pir gene completely eliminates the virulence of VPAHPND.6,11,18 VPAHPND secrete a binary toxin of PirABVP resulting in sloughing of tubule epithelial cells, and and acute dysfunction of the hepatopancreas.6,11 In the study,18 they reported that the binary toxin, PirABVP, triggers cell death via, which is stably inherited through a post-segregationally killing system that allows stable inheritance and is spread via a mechanism of conjugative transfer.18 Together with PirA, PirB combine to form the PirABVP complex, plays a critical role in intensifying the infection process, resulting in severe destruction of hepatopancreatic tissue in shrimp. It was reported that PirABVP enhances the pathogenicity of Vibrio parahaemolyticus strains, contributing to high mortality rates in affected shrimp populations.19,20 Even though binary toxin PirABVP is implicated in being a primary toxin causing AHPND, evidence indicated that toxin PirBVP is likely the main toxin protein enough driving the development of the disease pathogeny.3
Currently, many studies on AHPND have focused on identifying AHPND-causing strains. The characterization of the pathogenicity of various Vibrio parahaemolyticus strains are based on the structure of PirA and PirB. The representative genetic characteristics of the plasmid DNA containing the binary toxin genes (pirA and pirB) in Vibrio parahaemolyticus mutant strains R13, R14, XN87, as well as the reference strain A3, compared to non-pathogenic strain of A2, are summarized in Table 1.2,21–23 Based on these classification theme, V. parahaemolyticus carries both toxin genes causing AHPND are classified as Type I, while those with only one toxin gene that do not cause AHPND are Type II. The R13 mutant falls into the Type II category. However, the R14 strain, containing both PirA and PirB genes without causing AHPND, does not fit either Type I or Type II.
Roles of PirAVP, pirBVP, and pirABVP in AHPND
As mentioned above, PirAVP and PirBVP have been identified as the major factors responsible for inducing severe damage to hepatopancreatic tissues in shrimp. The releasing toxins, produced by Vibrio parahaemolyticus, play pivotal roles in the advancement of the disease since they can damage cellular integrity and provoke tissue necrosis. Therefore, understanding their mechanism of action is crucial for implementing targeted approaches to mitigate the impact of AHPND on shrimp aquaculture.
A prior report of structure analysis revealed that the tertiary structure of PirAVP, and PirBVP are closely resemble to Cry insecticidal toxin protein of Bacillus thuringiensis, highlighting a strong relationship between the PirAVP and PirBVP toxins and the Cry protein.18,24 Cry toxin protein comprised of three functional domains, including the pore-forming domain I – the pore-formation domain of α-PFTs colicin A, the receptor-binding domain II and the sugar-binding domain III.25 Referring to domain’s function, are suggested to play distinct roles: domain I is involved in forming the pore during membrane penetration, domain II facilitates receptor binding, and domain III is responsible for receptor recognition and membrane insertion.25,26 The resemblances of structure between PirAVP, PirBVP, and the Cry toxin, emphasizing their functional domains and cytotoxic mechanisms, are shown in Table 2. The resemblance of PirAVP and PirBVP to the Cry provides a basis for speculating on their functional roles, enabling their relevant to the toxin activities. PirAVP corresponds to the domain III of the Cry protein, while PirBVP has similar topology to The Cry domain I, and II. These similar topologies reveal that the binary toxin PirABVP has the Cry protein-like features, including three functional domains associated with cytotoxic activity, receptor binding, and membrane insertion.24 PirAVP has been reported to bind specially to ligands, such as monosaccharides like N-acetylgalactosamine (GalNAc) and oligosaccharides, located on the cell membrane, facilitating target-specific recognition. On the other hand, PirBVP, which consists of the N-terminal domain (PirBVPN) and the C-terminal domain (PirBVPC), induces cell death by forming pores in the host cell membrane and plays a role in protein–protein and protein–ligand interactions. The binding of PirABVP toxin and the early stages of pathogenesis in the digestive tracts of brine shrimp larvae.6 In shrimp aquaculture, the virulent bacteria responsible for AHPND, which carry the pVA1 plasmid encoding the PirABVP toxin genes, are absent in all non-AHPND bacterial species.
Albeit PirABVP toxin have been confirmed as the key factor causing the damage to hepatocellular of shrimps, resulting in shrimp mortality, many studies have proposed that PirBVP alone has the ability to cause cell damage.3,11,17,18 A study11 reported that after infected by AHPND-causing Vibrio parahaemolyticus, initial colonization occurred in the stomach, where the bacteria began producing the binary PirABVP toxin. The presence of the protein of PirAVP, and PirBVP were identified in the diseased shrimp’s hepatopancreas, they were also found in stomach, and hemolymph. The presence of PirBVP was earlier than PirAVP. Giving evidence, within 6 hours post-infection, the toxin of PirBVP was detected in the hepatopancreas, coinciding with the onset of characteristic histopathological signs of AHPND, such as sloughing of epithelial cells in hepatopancreas. The high concentration of Vibrio parahaemolyticus, and PirAVP were detected in the hepatopancreatic cells at later stages, 12 hours post-infection, and 18 hours post-infection, respectively.11 Given that PirBVP was enough to cause the AHPND in shrimps, and more toxic than PirAVP. The role of PirAVP was seemly to play an auxiliary role in the pathogenesis of AHPND, combining to PirBVP to form the full toxic of AHPND in shrimps.
Additionally, the toxin of PirBVP has been identified to first enter the stomach before being transferred to the hepatopancreatic cells, resulting in cell sloughing and cell death.11,27 Interestingly, our current data shows that the PirB toxin could be found in hemocytes, which suggest that it might have been transported there or to other tissues from the gastrointestinal system via hemolymph. A model of apoptosis in shrimp hemocytes induced by PirBVP was proposed.19 The release of PirBVP into the space of intratubular was facilitated by the disruption of hepatopancreatic tubules, then, transported into the hemolymph, resulting in targeting hemocytes. As the result, the PirBVP interacted with histone proteins, leading to the dephosphorylation of histone H3, therefore, triggering apoptosis-related pathways on hemocytes. It was suggested that underscoring the key role of PirBVP toxin.
The AHPND-causing pVA1, and gene of PirABVP can be transferred by gene transposition
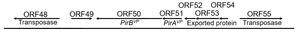
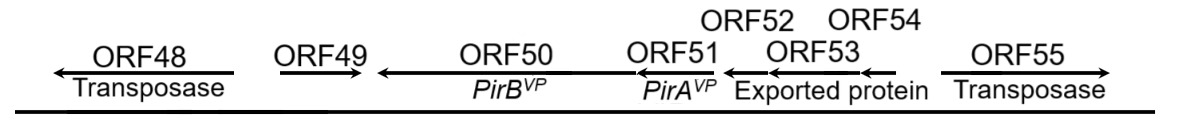
The PirABVP genes have been demonstrated to possess the capability for a horizontal gene transfer between organisms through gene transposition or homologous recombination.17,18 The horizontal gene transfer of PirABVP genes played a pivotal role in transformation of non-AHPND Vibrio parahaemolyticus into virulent strains of Vibrio parahaemolyticus.23 The transfer of of PirABVP can be illustrated through many key aspects. In the pVA1 (Accession number: KP324996), certain open reading frames, including ORF15, ORF48, ORF55, ORF57, and ORF68, known as transposase, were implicated in the virulence associated with this transformation.17 Three ORFs: ORF15, ORF48, and ORF55 shared 100% sequence identity, and being as members of the IS5 family of DDE transposases. These transposases carried the conserved DDE (Asp-Asp-Glu) motif, was essential for their roles in catalyzing DNA cleavage and strand transfer during transposition. ORF57, a member of the IS91 transposase family, contains a zinc-binding domain that likely contributes to DNA recognition and structural stability during transposition, although it is not a complete transposase. ORF68, from the Rpn family, is associated with recombination and promotes the integration of transposons into new genomic contexts. The cluster gene included PirAVP, and PirBVP was flanked by two transposase genes in opposite directions (ORF48 and ORF55) with inverted repeats in their terminals, called the pirAB-Tn903, or Tn6264, composite transposon (Fig. 1).17,28 Therefore, these structures enabled their excision, transfer, and integration into different plasmids or genomic locations. Regarding to the strain of XN87, a 1,053-bp transposase was inserted in PirAVP, resulting the disruption of its coding sequence. Consequently, the downstream of PirBVP gene expression was also affected. Such disruptions demonstrated the role of transposase-mediated events in the genetic variability and pathogenic potential of the pVA1 plasmid – causing the AHPND. The PirBVP gene, therefore, not only represented a pivotal factor in the pathogenicity of AHPND but also highlighted the role of horizontal gene transfer in the evolution and dissemination of bacterial virulence.
A novel Trb-Type IV secret system (T4SS) was responsible for mediating the conjugative transfer of the pVA1 plasmid across Vibrio spp. Total of 14 annotated genes, including traF, traG, and a cluster of 12 trb-genes (trbB1, trbB2, trbC, trbD, trbE, trbF, trbG, trbH, trbI, trbJ, trbL, and trbN) were identified to be components constituting a T4SS. Among them, trbE encoding ATPase, and traG encoding Type IV coupling protein (T4CP) were characterized as essential genes for the T4SS, as their deletion abolished the conjugative transfer of a pVA1-type plasmid from AHPND-causing Vibrio parahaemolyticus to Vibrio campbellii, while complementation restored it.29 The process of conjugation, a mechanism by which one genetic material transferred to a recipient bacterium, was processed by DNA transfer and replication system, while the coupling protein (CP) directs the protein-DNA complex through T4SS for secretion via pili, which are produced by the mating pair formation system (Mpf).17 The pVA1 plasmid contained most Mpf system genes except trbJ and trbK, with ORF11 annotated as a VirB1-like gene, essential for T4SS formation.17,30 This mechanism allows pVA1-related plasmids to transfer between bacterial cells. Therefore, these findings explored the novel ideas of the T4SS role in the horizontal transfer of virulence genes controlling the spread of virulence plasmid among Vibrio species, contributing to the pathogen diversity of AHPND.
Horizontal Gene Transfer of PirAVP, PirBVP, and PirABVP Toxins Converts Non-Vibrio Strains into AHPND-Causing Pathogens
The aforementioned PirABVP toxins, have structural and functional parallels with Pir toxin, cause the extensive tissue damage in the hepatopancreas of infected shrimp. Recent findings. have revealed that the gene of PirABVP are not exclusive to V. parahaemolyticus and may spread to other species via the horizontal gene transfer (HGT), thereby greatly expanding the range of potential AHPND-causing pathogens.31 According to the study of Muthykrishnan et al. (2019), they demonstrated that conjugatively transferring the binary genes of PirA, and PirB from a pathogenic V. parahaemolyticus to a non-Vibrio and non-pathogenic bacterium, identified as Algoriphagus sp. strain NBP, successfully transformed a non-pathogenic, non-Vibrio into disease-causing strain through HGT. HGT could occur between a pathogenic Vibiro and a non-Vibrio strain, particularly through conjugative plasmids, which are key vectors in the dissemination of virulence genes. The study further reported that the conjugative transfer of the pirAB-carrying plasmid from V. parahaemolyticus to Algoriphagus sp. strain NBP occurred with a high efficiency, ranging from 80% to 91% at incubation temperatures of 30°C and 40°C, developed acute hepatopancreatic necrosis disease (AHPND), as confirmed by histopathological analysis. Their study outcomes postulated that the presence of multiple non-Vibrio bacterial species in shrimp pond environments may harbor the PirA and PirB genes through horizontal gene transfer. It meant that horizontal gene transfer plays a central role in the emergence of new pathogenic lineages in aquaculture environments. These findings underscore the importance of broadening AHPND screening strategies to encompass non-Vibrio bacteria that may act as emerging reservoirs of pathogenicity.
Remarks and conclusions
The current review underscored the critical role of PirABVP the Acute hepatopancreatic necrosis disease caused by Vibrio parahaemolyticus, emphasizing their structural and functional resemblance to Cry insecticidal toxins. Moreover, the horizontal transfer of pVA1 plasmid carrying PirABVP gene further complicated disease control, underscoring the need for genetic monitoring in aquaculture. These highlighting provided the scientific foundation for developing targeted diagnostics, therapeutic interventions, and preventive measures. Expanding future research include non-Vibrio pathogens in AHPND screening may enhance disease management strategies and support the creation of sustainable solutions for shrimp aquaculture.
Acknowledgments
We gratefully thank AmphaOnco Pharmaceutical Corp, Ho Chi Minh City, Vietnam, for their financial support.
Authors’ Contribution
Conceptualization: Thuan D. Lao (Equal), Hoang A. Tran (Equal), Thuy A.H. Le (Equal). Formal Analysis: Thuan D. Lao (Equal), Linh T.T. Le (Equal), Nguyen H. Nguyen (Equal), Uyen P. Le (Equal), Hoang A. Tran (Equal), Phuong V. Nguyen (Equal). Investigation: Thuan D. Lao (Equal), Linh T.T. Le (Equal), Nguyen H. Nguyen (Equal), Uyen P. Le (Equal), Hoang A. Tran (Equal), Phuong V. Nguyen (Equal). Validation: Thuan D. Lao (Equal), Linh T.T. Le (Equal), Thuy A.H. Le (Equal). Writing – original draft: Thuan D. Lao (Lead). Writing – review & editing: Thuan D. Lao (Equal), Thuy A.H. Le (Equal).
Declaration of conflicting interests
The authors declare no potential conflict of interest with respect to the research, authorship, and publication of this article.
Informed Consent Statement
All authors and institutions have confirmed this manuscript for publication.
Data Availability Statement
All are available upon reasonable request.