Introduction
Nonspecific cytotoxic cells (NCC), also known as the fifth type of cytotoxic cells, are widely present in the lymphoid organs and peripheral blood of bony fish.1 The function of NCC cells is similar to that of Natural killer cells (NK cells) in mammals. NCC cells can recognize and destroy a variety of mutant cells, such as tumor cells, protozoa and virus-transformed cells, and play an important role in the host’s innate immunity.2 The activation of NCC is through the combination of the antigenic determinant of the target cell with the NCC receptor of the multimeric repeat sequence.3,4
Nonspecific cytotoxic cell receptor protein 1 (NCCRP1) is an important cell receptor on the cell membrane of NCC. This kind of receptor is a type I transmembrane protein with three important functional domains: antigen binding motif, signal transduction domain and transcription activation domain. The C-terminal of NCCRP-1 contains a region rich in tyrosine (Y), serine (S), and threonine (T), which are potential phosphorylation sites of protein kinases, and the N- terminal is also inside the cell, which contains a sequence rich in labeled amino acids, which is a sign of functional correlation.5,6 The typical proline-rich motifs (PRMs) in the N-terminal cytoplasmic region of NCCRP-1 are the characteristics of cytokine/hormone receptors, and can be used as “docking site” of JAK kinases to mediate the combination of them.5 NCCRP-1 interacts with protein tyrosine kinase, translocating the signal transducer and activator of transcription 6 (STAT6) into the nucleus and initiating transcription.5,7,8 NCCRP1 can induce cytotoxicity by recognizing and binding natural killer target antigen (NK tag) of various target cells, and may also activate JAK/STAT signal transduction pathway.5,9–12 NCCRP1 may play a dual role in the activation of NCC: (1) it acts as an antigen recognition molecule in the process of target cell lysis, and (2) it is the initiator of the release of cytokines by NCC.8,13,14
The amino acid sequence of NCCRP1 was initially deduced from the cDNA sequence of channel catfish NCCRP1.3 The determination of the cDNA sequence of NCCRP1 revealed that this conserved protein does not belong to the mammalian supergene family. Currently, NCCRP1 gene has been reported in various fish species.2–4,9,15,16 The NCCRP1 gene of Atlantic cod (Gadus morhua L.) is relatively small, which is different from the intron/exon structure of NCCRP1 of common carp and channel catfish, but it is also a relatively conservative gene.15 The transcriptional levels of NCCRP1 in Cyprinus carpio, Sparus aurata and Gadus morhua were all up-regulated to varying degrees after being stimulated by pathogens, indicating that NCCRP1 in fish might be involved in the immune response after pathogen infection.4,9,15
Megalobrama amblycephala is one of the important freshwater economic fish species in China. It is widely popular in the market because of its small head, tender meat and delicious taste. In recent years, with the rapid development of Megalobrama amblycephala breeding industry, pathogenic factors such as bacteria, viruses and parasites have caused rampant breeding diseases, and high-density breeding has led to eutrophication of water bodies, which has seriously threatened Megalobrama amblycephala breeding industry. The infection of Aeromonas hydrophila is one of the main causes of mass death of Megalobrama amblycephala. Its short infection time and high morbidity and mortality rate have become an important limiting factor for efficient green and healthy culture of Megalobrama amblycephala.17 NCCRP1 may play an important role in the immune defense of Megalobrama amblycephala against the infection of pathogenic bacteria such as Aeromonas hydrophila. However, up to now, the related research of NCCRP1 gene in Megalobrama amblycephala has not been reported, and the expression change law of NCCRP1 gene in different tissues after Aeromonas hydrophila infection is unclear. Therefore, in this study, the NCCRP1 gene of Megalobrama amblycephala was cloned for the first time, and its gene structure was analyzed to predict the function of NCCRP1. The expression of NCCRP1 gene in different tissues of Megalobrama amblycephala were detected by quantitative real-time PCR (qRT-PCR). After infected with Aeromonas hydrophila by immersion, the expression of NCCRP1 gene in different tissues of Megalobrama amblycephala was detected, so as to provide basic experimental data for the research on the prevention and control of bacterial diseases of Megalobrama amblycephala and its anti-infection mechanism.
Materials and Methods
Experimental materials
Megalobrama amblycephala (50±2 g) was taken from Hunan University of Arts and Science aquatic breeding base. Before sampling, Megalobrama amblycephala was temporarily cultured in the circulating water culture system of Hunan University of Arts and Science campus aquatic training base for two weeks. The radius and height of round plastic culture tank were 1m and 1.5m, respectively. During the culture process, the dissolved oxygen in the water was kept at 6.8±0.2 mg/L and the water temperature was kept at 26±2 °C. Carp adult fish extruded feed (Guangdong Tongwei Feed Co., LTD) which contained 31% protein was fed to the experimental fish at 2% of their body weight twice a day.
15 fish were sampled. After being anesthetized by MS-222 and taking blood from the tail vein, 100 mg of head kidney, liver, spleen, intestine, gill, posterior kidney and heart were taken from healthy Megalobrama amblycephala respectively, quickly freeze them with liquid nitrogen, and store in the refrigerator at -80 °C for later use. All procedures were performed in accordance with the “Guidelines for Experimental Animals” of the Ministry of Science and Technology (Beijing, China) and were approved by the Institutional Animal Care and Use Committee of the Hunan University of Arts and Science (number: JSDX-2022-001, date: March 2, 2022).
Aeromonas hydrophila infection and sample collection
Aeromonas hydrophila was isolated from diseased Megalobrama amblycephala. The monoclonal strain of Aeromonas hydrophila was selected and inoculated into Luria broth (LB) liquid medium. It was incubated overnight in a constant temperature shaker at 28 °C and 180 rpm, and then centrifuged at 4500 rpm for 8 minutes. The supernatant was discarded. Then diluted it with sterile PBS solution, the concentration of the diluted bacterial solution was detected by the plate live bacteria count method.
The concentration of Aeromonas hydrophila was 5.2×109 CFU/mL in the original culture. In the pre-experiment, Aeromonas hydrophila was diluted into five concentration gradients, with concentrations of 1.0×105 CFU/mL, 1.0×106 CFU/mL, 1.0×10⁷ CFU/mL, 1.0×108 CFU/mL and 1.0×109 CFU/mL. In each concentration group, Megalobrama amblycephala was immersed and infected with Aeromonas hydrophila for 30 minutes, and then transferred to aerated breeding buckets. The control group of Megalobrama amblycephala was immersed in sterilized water for 30 minutes, and then transferred to aerated breeding buckets. Each concentration group was set in three parallel repeated tanks with 30 M. amblycephala in each tank. The cumulative mortality rates for 96 hours after infection were counted. The mortality rates of the control group and the groups with different concentrations of Aeromonas hydrophila were 0, 7%, 16%, 21%, 27% and 35%, respectively. Through pre-experiment exploration, the concentration (1.0×109 CFU/mL) with the highest mortality rate was selected as the infection concentration for this experiment, which is in line with our previous study18
Megalobrama amblycephala of experimental group was immersed and infected with Aeromonas hydrophila at a concentration of 1.0×109 CFU/mL for 30 minutes, and then transferred to aerated breeding buckets. Three repeated breeding buckets were set up in the experimental group, with 30 Megalobrama amblycephala in each bucket. Samples were taken at 1 day, 3 day, 5 day, 7 day, 14 day and 21 day after infection, respectively. Untreated, normal and healthy Megalobrama amblycephala were selected as the control group. Fifteen Megalobrama amblycephala were taken from each of the control and experimental group at each time point. 100 mg of gills, liver, intestine and head kidney were taken respectively. After rapid freezing in liquid nitrogen, they were stored in a -80 °C refrigerator for future use.
Total RNA extraction and reverse transcription
The tissues of every five fish were mixed as a sample. Fifteen fish were sampled in each group and there were three parallel samples. Total RNA was extracted from the tissues of Megalobrama amblycephala using the RNA extraction kit (Simgen) in accordance with the operation instructions of the kit. The purity (OD260/OD280) and concentration of RNA were measured by a nucleic acid concentration meter. The integrity of RNA was detected by agarose gel electrophoresis. Non-degraded RNA was selected as a template, and reverse transcription was carried out by using Takara reverse transcription kit (Takara, Japan) according to the kit instructions. Remove genomic DNA reactions contained 2 μL 5×gDNA eraser buffer, 1 μL gDNA eraser, 1 μL total RNA, 1 μL rNase free dH2O, and the reaction condition was 42 °C for 2 min, termination at 4 °C. Reverse transcription reaction contained 10 μL primescript RT enzyme mix I, 1 μL primescript RT enzyme mix I, 1 μL RT primer mix which contained random 6 mers and oligo dT primer, 4 μL 5×primescript buffer 2, 4 μL RNase free dH2O, and reaction condition was 37 °C for 15 min, 85 °C for 5 sec, termination at 4 °C. The obtained cDNA template was stored in the refrigerator at -20℃.
Amplification of the cDNA sequence of the NCCRP1 gene
According to the cDNA sequence of NCCRP1 gene of cyprinid fish known in NCBI, primers (Table 1) were designed by Premier 5.0 software. The amplification of the NCCRP1 gene was carried out using the cDNA of the head kidney of Megalobrama amblycephala as the template. The PCR reaction conditions were: 94 °C for 5 minutes; 94 °C, 30 seconds, 56 °C, 30 seconds, 72 °C, 1 minute, cycle 30 times; 72 °C, 10 minutes, terminated at 4 °C. The PCR products were purified by DNA purification kit, linked with PMD19-T vector, transformed into DH5a competent cells, identified by bacterial liquid PCR, and positive clones were selected and sent to Hunan Qingke for sequencing.
Sequence analysis
In NCBI (http://www.ncbi.nlm.nih.gov/gorf/gorf.html), the ORF of NCCRP1 gene of Megalobrama amblycephala was searched and the amino acid sequence was predicted by open reading frame finder online software. The predicted amino acid sequence of NCCRP1 gene of Megalobrama amblycephala was analyzed and homologous sequence searched by online software blastp in GenBank. The amino acid domain of Megalobrama amblycephala NCCRP1 was labeled using SMART (http://smart.embl-heidelberg.de) and UniProt (http://www.uniprot.org/). The tertiary structure of Megalobrama amblycephala NCCRP1 was predicted by online software SWISS-MODEL (https://swissmodel.expasy.org). Multiple sequence alignment analysis was performed using Clustal X 1.83 software, and the genetic phylogenetic tree of the NCCRP1 gene was constructed using MEGA 5.1 software (NJ method, 1000 times).
Detection of NCCRP1 gene expression in Megalobrama amblycephala by qRT-PCR
Based on the cDNA sequence of the NCCRP1 gene of Megalobrama amblycephala, qRT-PCR primers (NCCRP1-QF and NCCRP1-QR) were designed using Premier 5.0 software. The qRT-PCR was carried out in triplicate on a Rotor-Gene Q real-time PCR Detection System (QIAGEN, Dusseldorf, Germany). With the β-actin gene as the internal reference gene, the relative expression levels of the NCCRP1 gene in various tissues of Megalobrama amblycephala were detected according to the operation instructions of the qRT-PCR kit (TaKaRa). The concentrations of different tissues cDNA was diluted into 50ng/μL. The total reaction volume of 20 μL contained 10 μL SYBR qPCR Mix, 1μL of each primer (10 mM), 2 μL cDNA and 6 μL ddH2O. qRT-PCR amplification conditions were 95°C for 5 min; 95°C, 20 s, 56℃, 20 s, 72℃, 20 s, 40 cycles; 72℃ for 8 min , terminated at 4 °C. The relative expression levels of the NCCRP1 gene in various tissues of Megalobrama amblycephala were calculated by 2-ΔΔCT method, as well as the expression change patterns of the NCCRP1 gene at different sampling time points after infection with Aeromonas hydrophila.
Data analysis
Experimental data are expressed as mean ± standard error. The expression differences of the NCCRP1 gene of Megalobrama amblycephala in various tissues and at different sampling time points after infection with Aeromonas hydrophila were analyzed by one-way ANOVA using SPSS19.0 software. P value < 0.05 indicated a significant difference, and P value < 0.01 indicated an extremely significant difference.
Results
Coding DNA Sequence of Megalobrama amblycephala NCCRP1 gene
The cDNA sequence of Megalobrama amblycephala NCCRP1 gene (GenBank Accession No: PX851883) was cloned by PCR technology. Its length was 726 bp, the ORF of Megalobrama amblycephala NCCRP1 was 702 bp, which encoded 233 amino acids. Its relative molecular weight and theoretical isoelectric point were 26.87KDa and 6.59, respectively. The amino acid sequence formula of Megalobrama amblycephala NCCRP1 was C1231H1849N323O348S4, which contained 29 acidic amino acids (Asp+Glu) and 27 basic amino acids (Arg+Lys). The results of amino acid analysis showed that Megalobrama amblycephala NCCRP1 had many functionally related sites, such as N- glycosylation site (NHSV, 51-54 aa; NFSG, 79-82 aa; NYSH, 181-184 aa), phosphorylation site of protein kinase C (TWK, 185-187aa; TSK, 231-233 aa), N-myristoylation site (GINHSV, 49-54aa; GNFSGW, 78-83 aa; GIPPGV, 96-101 aa), phosphorylation site of cyclic adenosine monophosphate and cyclic guanosine monophosphate-dependent protein kinase (KKVS, 187-190 aa), phosphorylation site of tyrosine protein kinase II (SVYE, 30-33aa; TSTE, 85-88 aa), tyrosine kinase phosphorylation site (KTSTEVLPY, 84-92 aa) (Figure 1).
Multiple comparison analysis of amino acid sequence of NCCRP1
In GenBank, the amino acid sequence of NCCRP1 gene of Megalobrama amblycephala was searched by Blastp online software, and compared with other fish NCCRP1. The similarity was as follows: Labeo rohita (87.12%), Carassius auratus (85.84%), Danio rerio (83.26%) and Cyprinus carpio (82.05%) (Table 2). Multiple sequence comparison and analysis of the amino acid sequence of Megalobrama amblycephala NCCRP1 with those of other species revealed a conserved antigen-binding motif of “xLPxxxWFxxEQxVxL” in the amino acid sequence of NCCRP1 of Megalobrama amblycephala (Figure 2). Meanwhile, the NCCRP1 of Megalobrama amblycephala has a conserved F-box-associated domain (FBA) and a proline-rich motif, and the N-terminal proline-rich motif, named “box-1” (pPXXXP) and “box-2” (pPXP), is responsible for the interaction and activation sequence of JAK (Figure 2).
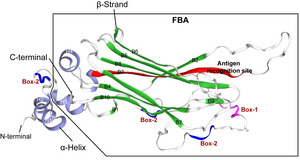
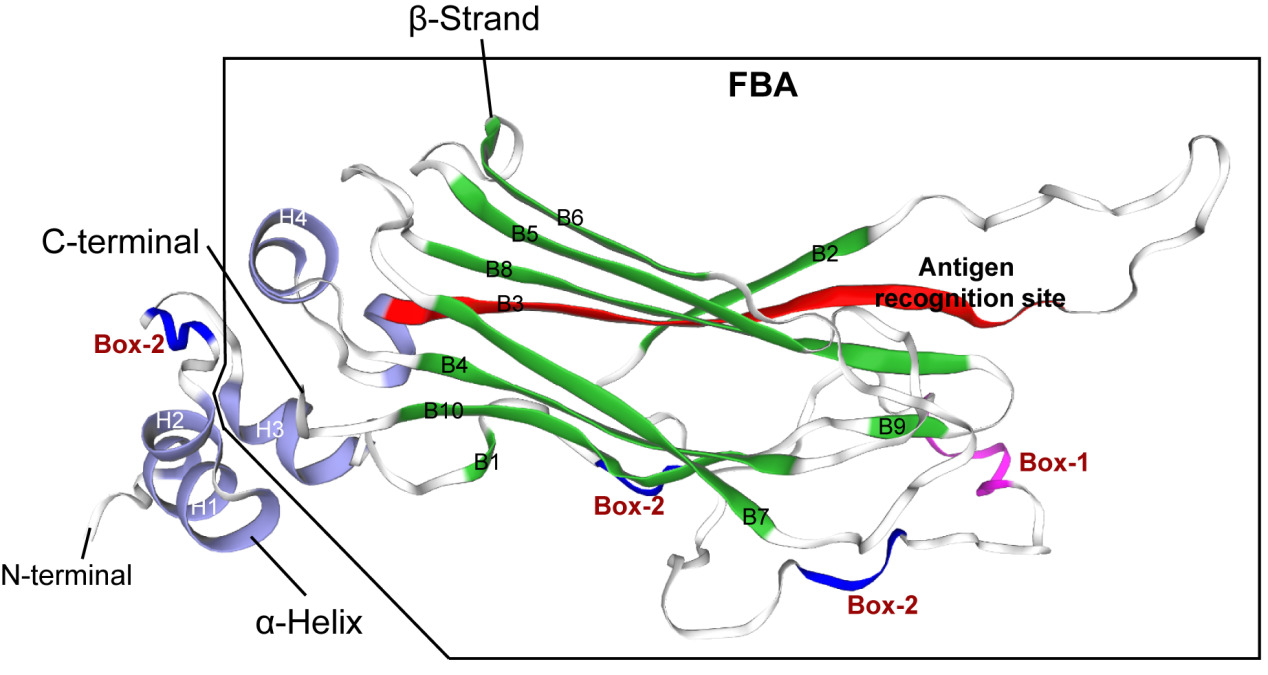
Prediction and analysis of the tertiary structure of NCCRP1 protein
The tertiary structure of Megalobrama amblycephala NCCRP1 was predicted by online software of SWISS-MODEL (https://swissmodel.expasy.org). It was found that the N-terminal of Megalobrama amblycephala NCCRP1 protein contained two α-helices (H1 and H2), and the C-terminal had a typical F-box domain, which was the binding site of substrate and showed a typical β-sandwich folding structure. The F-box-associated domain of Megalobrama amblycephala NCCRP1 was composed of 10 anti-parallel β-folded sheets (B1- B10) and two α-helices (H3 and H4) (Figure 3). In addition, the tertiary structure of Megalobrama amblycephala NCCRP1 showed the amino acid sequence at positions Leu106 to Ala122 (LPQHRWFSLEQHVDLKA), which was speculated to be the antigen recognition site of Megalobrama amblycephala NCCRP1 (Figure 3).
Phylogenetic tree of NCCRP1 gene
The amino acid sequence predicted by Megalobrama amblycephala NCCRP1 was subjected to multiple sequence alignment with the amino acid sequences of NCCRP1 of other species, and the genetic phylogenetic tree was constructed using the adjacency method (Figure 4).The NCCRP1 gene of Megalobrama amblycephala was clustered into one branch with that of Cyprinidae fish. The NCCRP1 of Megalobrama amblycephala had the closest genetic relationship with that of Ctenopharyngodon idellus.
NCCRP1 gene expression in different tissues of Megalobrama amblycephala
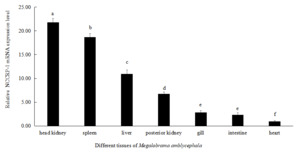
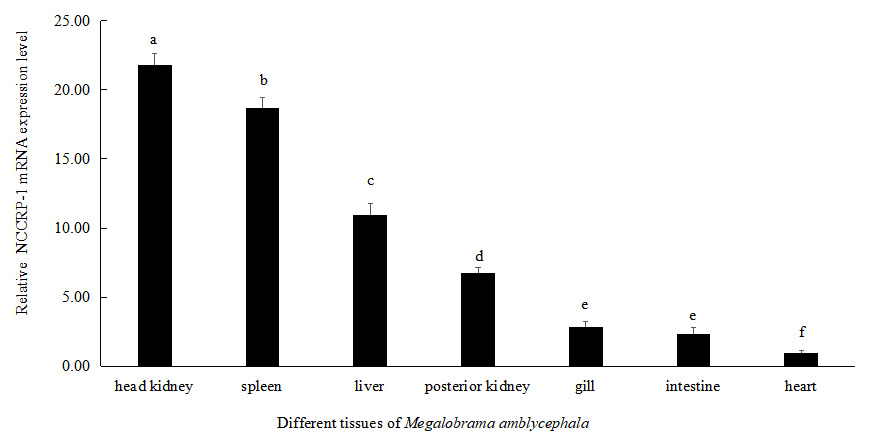
The expression level of NCCRP1 gene was highest in head kidney of Megalobrama amblycephala, followed by the spleen, liver and posterior kidney. There was no significant difference in the expression levels in intestine and gills of Megalobrama amblycephala (P > 0.05). The relative expression level of NCCRP1 gene was lowest in heart of Megalobrama amblycephala (Figure 5).
The immune response of the NCCRP1 gene after Megalobrama amblycephala infection with Aeromonas hydrophila
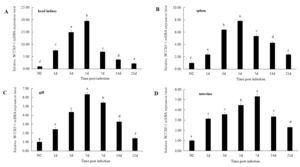
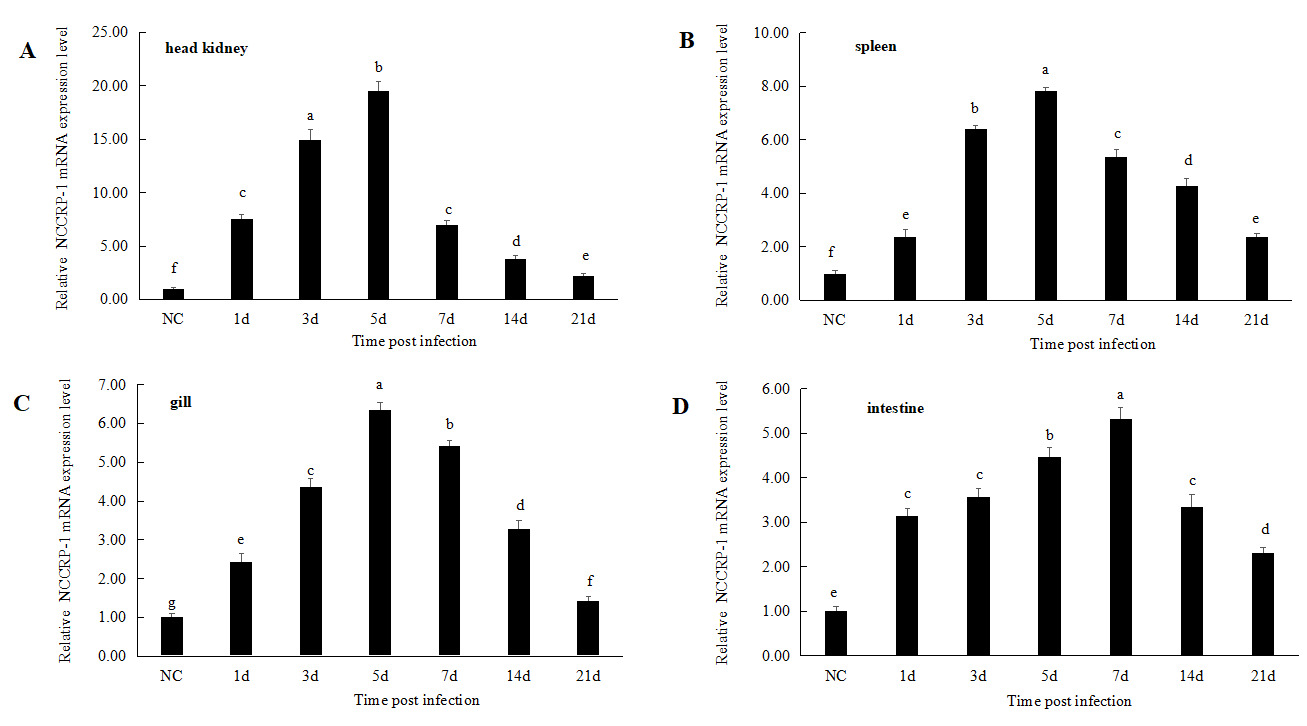
After infected with Aeromonas hydrophila, the relative expression levels of the NCCRP1 gene showed a trend of first increasing and then decreasing in the head kidney, spleen, gills and intestinal tissues during 21 days of infection (Figure 6). The relative expression level of the NCCRP1 gene in the head kidney significantly increased at 1 d after infection (P < 0.05, 7.58 fold), reached the peak at 5 d (19.48 fold), and then showed a downward trend, dropping to 2.24 fold that of the control group at 21 d. The expression level of the NCCRP1 gene in the spleen significantly increased at 1 d (P < 0.05, 2.37 fold), reached the peak at 5 d (7.82 fold), and gradually decreased at 21 d after infection. The expression level of NCCRP1 gene in the gill reached peak at 5 d after infection (6.35 fold), and then gradually decreased. It dropped to 1.42 fold that of the control group at 21 d, which was significantly higher than that of the control group (P < 0.05). The relative expression level of NCCRP1 gene in the intestine reached peak at 7 d after infection, which was 5.32 fold that of the control group, and decreased to 2.31 fold at 21 d after infection.
Discussion
NCC is regarded as a natural killer cell population similar to that of mammals, participating in the innate immune defense of bony fish. NCCRP1 is a key receptor protein on the NCC membrane, which is considered a specific marker of NCC.5,19 In the present study, the coding DNA sequence (CDS) of Megalobrama amblycephala NCCRP1 gene was cloned. It was deduced that Megalobrama amblycephala NCCRP1 contained 229 amino acids, and its relative molecular weight and theoretical isoelectric point were 26.87KDa and 6.59, respectively. The amino acid sequence deduced from Megalobrama amblycephala NCCRP1 gene was highly homologous to other reported fish NCCRP1, and the similarity with other fish NCCRP1 was more than 65%, which indicated that NCCRP1 was highly conserved in various fishes. Genetic phylogenetic evolution analysis shows that the NCCRP1 of Megalobrama amblycephala was clustered with that of Cypriniformes. Megalobrama amblycephala NCCRP1 and Ctenopharyngodon idellus NCCRP1 were clustered into a branch, and the parent relationship between Megalobrama amblycephala NCCRP1 and Ctenopharyngodon idellus NCCRP1 was the closest, which might be due to the fact that Megalobrama amblycephala belongs to cyprinidae fish, while Megalobrama amblycephala NCCRP1 was closely related to cyprinidae fish NCCRP1.
The amino acid sequence of Megalobrama amblycephala NCCRP1 contained three important functional domains: the N-terminal contains proline-rich transcriptional activation domains (Box-1 and Box-2), antigen-binding motility and the C-terminal signal transduction domain. Box-1 played a crucial role in the transcriptional activation of cytokines and hormone receptors mediated by the JAK/STAT pathway, and Box-2 might also bind to JAK kinases, thereby activating downstream cascade reactions.20–22 The protein interaction between NCCRP1 and JAK2 was reported in channel catfish, when NCC was activated by JAK2, STAT protein would be transferred to the nucleus, indicating that JAK/STAT pathway was involved in NCC activation.5 Moreover, the N-terminal PPP motif could also bind to SH3 of many phosphokinases, and the signal transduction domain of C-segment had been proved to play an important role in the activation of channel catfish NCC.5 Therefore, Megalobrama amblycephala NCCRP1 might have the similar functions in antigen recognition, NCC activation and JAK/STAT pathway regulation, while, its specific functions need further study.
The relative expression level of NCCRP1 gene was higher in the lymphoid organs, such as the thymus, head kidney, spleen and trunk kidney, which was lower in skin, muscle , liver, fat, heart and brain of Epinephelus coioides.6 The expression level of Channa argus NCCRP1 was highest in the liver and kidneys, followed by the spleen and peripheral blood, and the lowest in the brain and gills.23 The NCCRP1 expression were examined in all the tested tissues, which was highest in the liver of healthy Nile tilapia.24 The expression of Trachinotus ovatus NCCRP1 gene was detected in all tissues, which was the highest in the liver.25 The relatively high expression level of NCCRP1 gene was detected in the liver, spleen and head kidney of Lutjanus sanguineus.12 The expression of the NCCRP1 gene was various in different fish tissues. The NCCRP1 tends to be highly expressed in systemic immune organs across species. Expression in mucosal tissues varies widely. In this study, the NCCRP1 gene of Megalobrama amblycephala was expressed in all the tested tissues. The expression level of NCCRP1 gene was the higher in the systemic immune tissues than mucosal immune tissues of Megalobrama amblycephala. It indicated that the NCCRP1 gene might play an important role in the system immune tissue of Megalobrama amblycephala.
After Vibrio alginolyticus infection, the expression level of Lutjanus sanguineus NCCRP1 increased and reached peak at 4h.12 After infection with Cryptocaryon irritans, the mRNA level of NCCRP1 was up-regulated at 6h in skin of Epinephelus coioides, which was reached peak at 3 d post the primary infection.6 The expression patterns of NCCRP1 differed in the systemic immune organs of Epinephelus coioides, in the spleen, NCCRP1 expression was gradually increased in the early infection period and decreased sharply at 3d post the primary infection with Cryptocaryon irritans, whereas, the expression of NCCRP1 was depressed in the head kidney after infection.6 After Streptococcus iniae infection, the mRNA level of NCCRP1 gene was significantly increased in the head kidney, liver and intestine of Trachinotus ovatus at 24 h post-stimulation, which was significantly increased in the brain at 48 h post-stimulation (Huang et al., 2024). The expression of NCCRP1 gene was significantly up-regulated at 6 h and which remained contant in the spleen of Trachinotus ovatus after Streptococcus iniae infection.14 After Nocardia seriolae infection, the expression of Channa argus NCCRP1 gene was continuously up-regulated in spleen, and the mRNA level of NCCRP1 gene in kidney and liver reached the peak at 1 d and 2 d after infection, respectively.23 In this study, the relative expression levels of the NCCRP1 gene showed a trend of first increasing and then decreasing in the head kidney, spleen, gills and intestine of Megalobrama amblycephala during 21 days of infection with Aeromonas hydrophila. The expression level of the NCCRP1 gene in the head kidney, spleen and gills was reached the peak at 5 d, which was reached the peak at 7 d in the intestine after infection, indicating that the activation of immune response is a very complicated and time consuming process. Moreover, the relative expression level of the NCCRP1 gene at the peak in the head kidney and spleen was higher than that in the intestine and gills. The head kidney and spleen are the main enrichment areas of NCC, and they are the primary battlefields of Aeromonas hydrophila attack. A large number of NCCRP1 are needed to arm NCC cells in head kidney and spleen to realize rapid identification, killing and signal amplification. This is the functional basis of high expression of NCCRP1 in systemic immune tissues.
In this study, the CDS sequence of NCCRP1 gene of Megalobrama amblycephala was cloned by PCR, and its gene structure was analyzed. The expression of NCCRP1 gene was detected in different tissues, which was the highest in the head kidney of Megalobrama amblycephala. After infection with Aeromonas hydrophila, the expression of NCCRP1 gene was first increased and then decreased in immune-related tissues of Megalobrama amblycephala. NCCRP1 plays an important role in resisting Aeromonas hydrophila infection. However, the specific mechanism of its action remains to be further studied.
Acknowledgments
This work was supported by the Guangxi Natural Science Foundation (Grant No. 2022JJA130338), the National Natural Science Foundation of China (Grant No. 32360922), Hunan Natural Science Foundation (Grant No. 2025JJ50145) and the open Foundation of the State Key Laboratory of Development Biology of Freshwater Fish Sub-Center for Health Aquaculture (2025KF14).
Authors’ Contribution
Conceptualization: XinYue Zhao, Hu Xia; Methodology: XinYue Zhao, Yanan Gong, Liangguo Liu; Formal analysis and investigation: Hu Xia, Yanan Gong, Fuyan Chen, XinYue Zhao; Writing - original draft preparation: XinYue Zhao, Fuyan Chen, Jiahui Hu, Hu Xia; Writing - review and editing: Hu Xia, Liangguo Liu; Funding acquisition: Hu Xia; Resources: Rurou Fu; Supervision: Hu Xia.
Competing of Interest – COPE
The authors of this study declare no conflict of interests.
Ethical Conduct Approval – IACUC
The experiment was conducted with approval from the IACUC (Institutional Animal Care and Use Committees) of Hunan University of Arts and Science, Changde, China.
Informed Consent Statement
All authors and institutions have confirmed this manuscript for publication.
Data Availability Statement
All are available upon reasonable request.