Introduction
The Hippo signaling pathway plays a crucial role in regulating cell proliferation, apoptosis, and immunity in animals.1–4 Warts (Wts), which belongs to the nuclear Dbf2-related (NDR) protein kinase family,was first reported in Drosophila melanogaster.1 Wts, Salvador (Sav), Yki, and Hippo are key component in the Hippo signaling pathway which is highly conserved in Drosophila and mammals.5 When wts is mutated, cells undergo excessive growth and form tumors.2 Wts was phosphorylated and activated by Hippo.2 Yki was regulated by Wts through two Wts phosphorylation sites on Yki.2,6 After being phosphorylated, Yki was sequestered in the cytoplasm and could not regulate the transcription of downstream genes related cell proliferation in the nucleus.7
Currently, few reports about the function of Wts are available in aquatic animals.2 The large tumor suppressor (Lats) is a homologous gene of Wts in vertebrates and Crassostrea hongkongensis.5,8 Lats1/2 play an important role in the formation of gastrula in Danio rerio.9 When lats1/2 were knocked out, the number of cardiomyocyte was increased in atria during early development of D. rerio.10 Lats1/2 could phosphorylate angiomotin, which inhibited endothelial cell migration in vitro and angiogenesis in zebrafish embryos in vivo.11 Lats was highly expressed in haemocytes and triggered hemocytic apoptosis in C. hongkongensis, which suggested that Hippo signaling pathway was involved in the immune defense process of oyster.8 When Hippo and Wts were silenced, the ability of hemocyte phagocytosis become weakened, which proved that the Hippo signaling pathway could positively regulate the cellular immunity in P. vannamei.12 The white spot syndrome virus (WSSV) in shrimp P. vannamei increased after Wts and Hippo were silenced in vivo, which indicated a positive role of Hippo signaling in antiviral response.13
Penaeus japonicus, also named Marsupenaeus japonicus, is widely distributed in the Indo-Western Pacific, including China, Japan, Southeast Asia, East Africa, and Australia.14 It is an economically important shrimp species in China.15 P. japonicus is cultured along southeast coastal areas of China due to its fast-growth and strong adaptability.14 The production of P. japonicus was 4.62 ten thousand tons in China in 2022.15
The shrimp aquaculture industry has been threatened by various pathogens including viruses, bacteria and fungi recently.16,17 Vibrio species are commonly found in marine and estuarine environments. Vibrio harveyi is thought as one of the main pathogens in shrimp, which cause several acute diseases such as hepatopancreatic necrosis, bacterial white-tail disease, black shell disease and light-vibriosis.17 Understanding the molecular regulatory mechanism of defense against bacteria in shrimp is useful to design strategy to prevent and control bacterial diseases. Nowadays, there are some reports on the molecular mechanism of innate immunity in P. japonicus.18 However, it remains unclear whether Wts is involved in the innate immunity of P. japonicus.
In this study, the cDNA and genomic DNA sequences of Wts of P. japonicus (named PjWts) were obtained, characterized and analyzed. The role of Wts in response to V. harveyi infection was investigated with the real-time quantitative polymerase chain reaction (RT-PCR) and RNA interference (RNAi) techniques. Our study provides a theoretical basis for further study of the role of Wts and the Hippo signaling pathway in the innate immunity of P. japonicus.
Materials and Methods
2.1. Experimental Animals and Sample Preparation
All of P. japonicus individuals were purchased from a shrimp aquaculture company in Fujian Province. Several tissues including the hepatopancreas, muscle, lymph, gills, ganglia, stomach, heart, hemolymph and eye stalks were separately collected from five shrimps (20.7 ± 1.2g). All of tissues were frozen with liquid nitrogen quickly, and stored in -80℃ until used.
A total of 410 healthy and active individuals of P. japonicus (31±2.1 g) were chosen randomly and cultured in several concrete ponds (20 m2) three days for the experiment of bacterial challenge. The salinity and temperature of seawater was 35‰and 25℃, respectively. Two-third seawater was changed and the residual feed/waste was removed everyday. All shrimps were fed a commercial mixed feed supplied by Haid Group Co., Ltd. (Guangdong, China).
2.2. Bacterial challenge
The shrimps were divided into two groups designated as control group (CG) and experimental group (EG). Each group was set up in triplicate (i.e., three biological replicates). V. harveyi was used for the bacterial stimulation experiment. Shrimps in the EG were intramuscularly injected with 50 μL of resuspended bacteria solution (1x 106 cfu/ml), while the CG were injected with 50 μL phosphate-buffered saline (PBS; 137 mM NaCl, 2.7 mM KCl, 8 mM Na2HPO4, 2 mM KH2PO4, pH 7.4).
The hepatopancreas and gills were collected from five shrimps at different time points (0, 3, 6, 12, 24, 48 and 72 h post-injection) for RNA extraction. All samples were frozen immediately with liquid nitrogen quickly, and then stored at -80°C until used.
2.3. RNA extraction and cDNA synthesis
Total RNA from each tissue was isolated using Trizol reagent (Invitrogen, USA) following the manufacturer’s instructions. The quality and concentration of RNA were examined by 1.5% agarose gel electrophoresis and Nanodrop 2000 spectrophotometer (Thermo Scientific, USA), respectively. A total of RNA 1 μg of total RNA was used as the template for reverse transcription reaction with a PrimeScriptTM RT reagent Kit with gDNA Eraser (Takara, Japan), following the manufacturer’s instructions.
2.4. Validation of Wts from Transcriptome Data
The full-length of cDNA sequence of Wts was obtained from transcriptomic sequences (SRX2030618) of P. japonicus. All primers were designed with Primer Premier 5 (Table 1). A pair of primers (Wts-S1/R1) was used to confirm the accuracy of Wts sequence. Polymerase chain reaction (PCR) was performed in a 20 µl reaction volume containing 12 µl ddH2O, 2.0 µl 10 × Ex Buffer,1.6 µl dNTP mixture (2.5 mmol/L), 1.6 µl MgCl2 (25 mM), 0.8 µl of each of primer (10 µM), 1 µl cDNA (50 ng/μL), and 0.2 µl Ex Taq DNA polymerase (5 U/µl) (TaKaRa, Japan). The PCR product was purified by SanPrep Column DNA Gel Extraction Kit (Sangon, Shanghai, China), and then sequenced directly by Sangon Biotech (Shanghai) Co., Ltd. (China).
2.5. Bioinformatics analysis
The open reading frame (ORF) and predicted protein of Wts were analyzed by ORF finder (https://www.ncbi.nlm.nih.gov/orffinder/). The physicochemical property of protein was analyzed using ProtParam (http://web.expasy.org/protparam/). The similarity of PjWts with Wts from another species was analyzed with Global Align (https://blast.ncbi.nlm.nih.gov/Blast.cgi?). The structure of PjWts protein was analyzed by Conserved domains (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). The signaling peptide was predicted by SignalP5.0 (http://www.cbs.dtu.dk/services/SignalP/). The transmembrane domain was predicted by TMHMM (https://services.healthtech.dtu.dk/service.php?TMHMM-2.0). Multiple alignment of protein was analyzed by Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/). The phylogenetic tree was constructed by MEGA 7.0 software and neighbor joining was used and branching reliability was tested by bootstrap resampling with 1000 pseudo-replicates.
2.6. Expression of Wts in different tissues
The relative expression levels of Wts in different tissues from five shrimp were analyzed using quantitative real-time PCR (qRT-PCR) with primers qWts-S and qWts-A (Table 1). All qRT-PCR assays were carried out in three biological replicates. The qRT-PCR was performed using SYBR Premix Ex Taq (Takara, China) according to the manufacturer’s instructions, and carried out on LightCycler® 480 II (Roche, Switzerland). The β-actin (GenBank accession No.: AB055975.1) gene was chosen as the reference gene.19 PCR programme was as follow: 95 °C for 30 s, 40 cycles of 95 °C for 5 s, 60 °C for 15 s, finally a melting curve was obtained at the end of the reaction and a single peak was observed. The acquired data were calculated using a 2–ΔΔCT method,20 and SPSS 19.0 software (IBM, USA). One-way analysis of variance (ANOVA) was used to evaluate significant differences (P < 0.05).
2.7. RNA interference (RNAi) and survival rate analysis
The recombinant plasmids, pD7-Wts and pD7-GFP, were established, respectively. The primers specific to target genes were dsWts-S1/A1 and dsGFP-F/R with T7 promoter sequence (Table 1). Sense and anti-sense DNA templates for transcription in vitro were produced by employing pD7-Wts and pD7-GFP recombinant plasmids as templates via PCR. dsRNA-Wts and dsRNA-GFP were synthesized in vitro with a Transcription T7 kit (TaKaRa Biotechnology) following manufacturer protocols. dsRNA-GFP was served as a control. The quality of the dsRNA was evaluated using 1.2% agarose gel electrophoresis. dsRNA was stored at −80 ˚C until used.
dsRNA targeting Wts (dsRNA-Wts) or green fluorescent protein (dsRNA-GFP; control) (50 μg per shrimp) was injected into the abdominal segment of each shrimp.21 The hepatopancreas and gills were scissored from four shrimps at 0h, 12h, 24h and 48h post-injection, then immediately frozen in liquid nitrogen. The expression of PjWts mRNA at different time points was checked by qRT-PCR as described in Sections 2.1 and 2.6. All samples were tested in triplicates.
To investigate the role of Wts during V. harveyi infection, shrimps were injected with dsRNA-Wts, dsRNA-GFP or PBS firstly, then challenged by V. harveyi (106 CFU.mL-1) 48h later. The number of dead shrimps in each groups was recorded within 96 h. Statistics were processed by GraphPad Prism 8.0 Software (GraphPad Software, USA).
Results
3.1. Cloning and sequence analysis of PjWts
The full-length cDNA sequence of PjWts was 4,123 bp in length, including an open reading frame (ORF) of 3,438 bp encoding 1,145 amino acids, a 5`untranslated region (5`UTR) of 225 bp and a 3`UTR 460 bp (Figure 1). The molecular weight and isoelectric point of PjWts protein were predicted to be 125.607 kDa and 8.96, respectively. No transmembrane domains or signal peptides were identified. Two conserved domains were detected: a Mob-binding domain (Mob, 677–752 aa) and a serine/threonine kinase catalytic domain of the large tumor suppressor family (STKc_LATS, 753–1115 aa). The PjWts cDNA sequence was submitted to GenBank with the accession number PP155416.1.
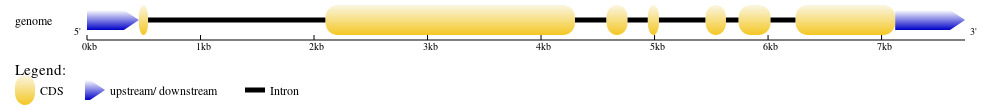
The genome sequence of PjWts was identified from genome sequence of P. japonicus (GenBank accession No.:NW_025035089.1). After alignment with the cDNA sequence of PjWts, seven exons and six introns were identified in the genomic sequence of PjWts. The longest and shortest exon was exon 2 (2,197bp) and exon 1 (81bp), respectively (Figure 2). The longest and shortest intron was intron 1 (1,564bp) and intron 5 (112bp) (Figure 2). The exon-intron junction regions conformed to the GT/AG rule. The genomic sequence of PjWts has been submitted to GenBank (Accession number: PP155417.1).
The protein sequences of PjWts shared over 90% identity with the Wts from other shrimp species, and low identity with LATS1/2 protein from vertebrates (Table 2). The similarity of PjWts with that in A. dorsata, E. sinensis, P. trituberculatus, C. quadricarinatus, H. americanus and D. melanogaster was 51%, 80%, 80%, 90%, 91% and 45%, respectively. The MobB and STKc_LATS domains were highly conserved in all test species. A total of 439 amino acids (aa) were detected in these two domains. From invertebrates to vertebrates, more than 249 completely conserved amino acids (accounted for 59%) and 91 relatively conservative amino acids (accounted for 20.78%) were detected in these two domains (Figure 3).
The phylogenetic tree of Wts and Lats1/2 from 19 species was constructed (Figure 4). This tree was divided into two main clades: one corresponding to vertebrates and the other to invertebrates. PjWts first clustered with Wts proteins from other Penaeus shrimp species, and then further clustered with those from other crustaceans. The LAST1 and LAST2 from Danio rerio, Homo sapiens and Mus musculus each formed separate subclusters within the vertebrate clade before merging into a single vertebrate-specific cluster.
3.2. Expression profiles of PjWts in different tissues
The Result of qRT-PCR showed that PjWts was expressed in all eleven examined tissues (Figure 5). The expression level of PjWts was the highest in the ganglia, followed by the gills, ovaries, and lymphoid organs. In contrast, relatively low expression levels of PjWts were detected in the eyestalks, hemolymph, and eyes (Figure 5).
3.3. Expression level of PjWts after bacterial challenge
The results of the PjWts mRNA expression in gills and hepatopancreas after V. harveyi infection are shown in Figure 6. The mRNA expression level of PjWts was significantly upregulated at 3 h post-infection. It reached peak levels at 6 h in the gills and 12 h in the hepatopancreas, respectively. At 72 h post-infection, the mRNA expression of PjWts was significantly higher than that in the control group (P < 0.05).
3.4. RNA interference efficiency and Survival rate
The effect of PjWts knockdown in hepatopancreas and gills was evaluated using RNAi technology (Figure 7). The expression levels of PjWts in the hepatopancreas and gills were significantly decreased at 12h after dsRNA-Wts injection (Figure 7). Compared with the control group, the interference effect was significant at 12 h, 24 h, and 48 h after dsRNA-Wts injection in the hepatopancreas, with knockdown percentages of 60.54%, 75.64%, and 97.30%, respectively. In the gills, the knockdown percentages were 54.87%, 75.55%, and 87.12%, respectively.These results indicate PjWts were significantly knocked down in hepatopancreas and gills.
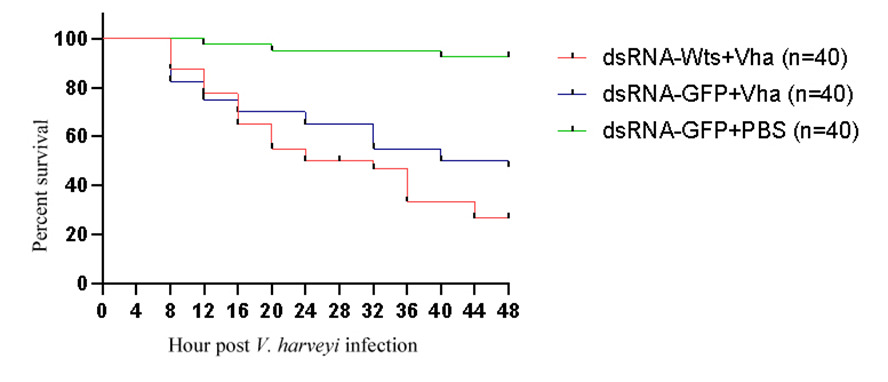
The survival rates of the shrimps in three groups were recorded after V. harveyi infection (Table 3). The survival rates of dsRNA-PjWts + V. harveyi, dsRNA-GFP + V. harveyi and dsRNA-GFP + PBS group were 20%, 47.5% and 92.5%, respectively (Figure 8, Table 3).
Discussion
The Hippo signaling pathway plays an important role in innate immunity of many animals such as D. rerio, C. hongkongensis and P. vannamei.2,8,12,13,22 The Hippo signaling pathway was involved in immune responses to bacteria and viruses in P. vannamei.12,13 However, no studies have reported on the Hippo signaling pathway in P. japonicus.
As a core component of the Hippo signaling pathway, Wts is essential for its function.4,22 The cDNA and genomic sequences of PjWts were isolated and characterized in this study. The ORF of PjWts was 3,438bp in length, encoded 1,145 putative amino acids (Figure 1), which was similar with the length of PcWts (1,137 amino acids) and LvWts (1,140 amino acids).
The identity of PjWts protein sequence with LATS from vertebrate was ranged from 38% to 41% (Table 2). In contrast, PjWts shares a high sequence identity (99%) with Wts proteins from other Penaeus species (Table 2). Both of MobB and STKc_LATS domain of Wts were detected, which maintain high conservation among various species in this study (Figure 2). These results suggest that the Wts protein, particularly its MobB and STKc_LATS domains, is evolutionarily conserved. This conservation implies that Wts may exert similar physiological functions across different species. Notably, STKc_LATS domain could phosphorylate protein and maintain cell activity23—a function that likely contributes to PjWts’ role in immune regulation.
The genomic sequence of PjWts consists of 7 exons and 6 introns, which is consisted with the structure of Lats1/2 genome in human and mice but differs from that of Lats1/2 in zebrafish (which contains 8 exons).9 This observation indicates that the genomic structure of Wts is highly conserved with minimal evolutionary variation.
Phylogenetic tree analysis revealed that P. japonicus clusters with other crustaceans—a result consistent with traditional taxonomic classifications (Figure 4). Meanwhile, LATS1 and LATS2 from vertebrates each form distinct subclusters within the vertebrate branch (Figure 4). To date, no Wts homologs have been reported in shrimp (beyond the present study). Evolutionarily, vertebrate LATS1 is thought to originate from invertebrate Wts, whereas LATS2 may have emerged via whole-genome duplication during evolution9—a process that could explain the functional divergence of LATS1/2 in vertebrates compared to invertebrate Wts.
PjWts mRNA is expressed in various tissues of P. japonicus (Figure 5), which is consistent with the tissue expression pattern of Wts in P. vannamei.13 These results suggest that Wts may perform multiple physiological functions in shrimp. Additional experiments are required to verify this hypothesis.
In mammals, the Hippo signaling pathway plays an important role in the development of the nervous system, including regulating the proliferation and differentiation of neural cells, and promoting the development of synapses, the corpus callosum, and the cerebral cortex (Li X, et al., 2021). Notably, the expression of PvWts in P. vannamei is highly abundant in ganglia13—a finding consistent with our observation of high PjWts expression in P. japonicus ganglia (Figure 5). This conservation in ganglionic expression implies that Wts may be involved in neurodevelopment in shrimp.
In mice, only a few follicles and no corpus luteum were observed in the ovaries of Lats1-knockout individuals, suggesting that LATS1 supports fertility in mammals.24 Tsoi et al.25 were the first to demonstrate that LATS1/2 play a critical role in maintaining the genetic program of ovarian granulosa cells. In the present study, we also detected high PjWts expression in the ovary of P. japonicus—a result consistent with the aforementioned mammalian studies (Tsoi et al., 1999; St John et al.24). However, the specific function of Wts in shrimp ovaries remains unclear, and further research is needed to elucidate its role in shrimp reproduction.
In shrimp, hemolymph, gills, and lymphoid organs play critical roles in innate immunity, including both cellular and humoral immune responses.26 For instance, Lats mRNA is highly expressed in the hemocytes and gills of C. hongkongensis.8 In the present study, we detected high PjWts expression in the gills and low expression in the hemolymph of P. japonicus (Figure 5)—a pattern consistent with the tissue expression of PvWts.13
Following V. harveyi challenge, the expression of PjWts was significantly upregulated, reaching peak levels at 6 h in the gills and 12 h in the hepatopancreas, respectively (Figure 6). This result suggests that PjWts may be involved in the defense response against Vibrio species, which aligns with findings from previous studies. For example, in C. hongkongensis, Lats expression in hemocytes was significantly increased at 3 h post-injection with V. coralliilyticus and reached a maximum at 6 h.8 In p. vannamei, PvWts expression was significantly upregulated at 12 h post-Vibrio infection and increased rapidly from 48 h to 72 h.12 Additionally, PvWts expression was elevated in P. vannamei following WSSV infection, accompanied by increased hemocyte activity.13
Previous studies have shown that animals infected with bacteria exhibit injury or mortality when Wts or Lats is knocked out or silenced. For example, the mortality rate of D. melanogaster infected with gram-positive bacteria increases following Wts knockout.2 In P. vannamei, silencing of PvWts enhances the infectivity of WSSV.13 Additionally, silencing Wts in shrimp reduces the phagocytic activity of hemocytes. Consistent with these findings, the mortality rate of shrimp infected with Vibrio exceeds 80%—a rate higher than that of uninfected shrimp.12
In mammals, knockout of Lats1 in mice leads to the development of soft tissue sarcomas and ovarian stromal cell tumors, indicating that Lats1 functions to inhibit tumorigenesis.24 In the present study, we also observed that the expression of PjWts in the gills and hepatopancreas gradually decreased within 48 hours after PjWts silencing, which was significantly different from the results in the control group (Figure 7). This finding confirms that PjWts was successfully inhibited.
Notably, the survival rate of P. japonicus infected with V. harveyi was only 16% following PjWts silencing. In contrast, the survival rate of P. japonicus infected with V. harveyi was 40% when PjWts was not silenced (Figure 8). These results suggest that PjWts may inhibit the virulence of V. harveyi in shrimp.
In summary, the cDNA and genomic DNA sequences of the PjWts gene were isolated and characterized in this study. The Mob and STKc_LATS domains of PjWts are highly conserved across various species. PjWts mRNA is highly expressed in the ganglia and gills of P. japonicus. Following V. harveyi injection, the mRNA expression of PjWts was significantly upregulated in the gills and hepatopancreas. After PjWts silencing, the survival rate of P. japonicus infected with V. harveyi was significantly lower than that of shrimp in the other groups. These results suggest that PjWts may be involved in the innate immunity of P. japonicus, laying a foundation for further investigations into the function of the Hippo signaling pathway in this species.
Acknowledgments
This work was supported by Open project of Key Lab of Freshwater Biodiversity Conservation, Ministry of Agriculture and Rural Affairs of China (LFBC1114); Open Program of Key Laboratory of Cultivation and High-value Utilization of Marine Organisms in Fujian Province(2025fjscq04); Hunan Natural Science Foundation Youth Project (2024JJ6331).
Authors’ Contribution
Chao Peng: Methodology, formal analysis and writing - original draft. Shuiqing Wu: Methodology, data curation. Haihong Huang: Methodology, data curation. Liye Shao: collected the samples and performed the bioinformatic. Yibin Yang: designed, conceived of the study, supervision. All authors gave final approval for publication.
Competing of Interest – COPE
No competing interests were disclosed.
Ethical Conduct Approval – IACUC
P. japonicus used in this experiment do not belong to vertebrates or regulated invertebrates, and are not endangered protected animals.
Informed Consent Statement
All authors and institutions have confirmed this manuscript for publication.
Data Availability Statement
All are available upon reasonable request.




_and_hepatopancreas(b)_after_*v._harveyi*_.png)
_and.png)


_and_hepatopancreas(b)_after_*v._harveyi*_.png)
_and.png)